Introduction
Banana shrimp Penaeus merguiensis is widely distributed in the western part of Indo-Pacific (i.e. from the Persian Gulf to Hongkong and Australia; Aziz et al., 2020; Wanna et al., 2005). Among penaeid species, P. merguiensis is regarded as a high value species owing to its meat quality and texture (Aziz et al., 2020; Prasertlux et al., 2024). Breeding programs of P. merguiensis had been initiated internationally between 1990s to early 2000s (Aziz et al., 2020). Consequently, its aquaculture production worldwide peaked in 2006 (at 96,633 tons) but it was consistently reduced to 24,681 tons in 2016 (FAO, 2019).
Thai Department of Fisheries has concerned about the sustainability of the shrimp industry. Currently, an introduced Pacific white shrimp Litopenaeus vannamei contributes over 95% of the annual aquaculture production (Prasertlux et al., 2024). However, P. merguiensis is a local species which is considered as a potential alternative species for shrimp aquaculture. In Thailand, neither semi-intensive nor intensive culture system of P. merguiensis is still not well developed and genetically improved broodstock and larvae are not commercially available. Therefore, selective breeding programs of local penaeid species like P. merguiensis should be implemented for the sustainability of the shrimp industry.
The information on genetic diversity and population differentiation is essential for implementation of appropriate resource management and the construction of effective breeding programs of economically important species (Sajeela et al., 2019; Vaseeharan et al., 2013). Analysis of genetic diversity using mtDNA polymorphism is straightforward. Unlike nuclear DNA, animal mtDNA is haploid and transmitted maternally. Accordingly, the effective population size estimated from mtDNA is smaller than that of nuclear markers (Birky et al., 1989; Khamnamtong et al., 2009). This results in the sensitivity to inbreeding and bottleneck effects relative to that of nuclear DNA markers.
Mitochondrial gene polymorphism, for example, 16S ribosomal DNA (16S rDNA), Cytochrome c oxidase subunit I (COI) and the control region has been used for various applications including species authentication, stock structure analysis, and genetic boundaries within species (Abdul Halim et al., 2021; Klinbunga et al., 2007; Mohammed-Geba & Yousif, 2022). In penaeid shrimp, COI is effectively applied for determination of intraspecific genetic variability and discrimination of cryptic species (Hualkasin et al., 2003; Sajeela et al., 2019). It is the most common gene region used for DNA barcoding of animals (Ward et al., 2005).
Previously, intraspecific genetic structure of P. merguiensis between Gulf of Thailand and the Andaman Sea were reported with limited sample size (n = 26) from 7 geographic locations composing of Trat, Surat Thani, Nakhon Si Thammarat and Songkhla located in the east of peninsular Thailand and Ranong, Satun and Phuket located in the west of peninsular Thailand (Hualkasin et al., 2003). Subsequently, three nuclear loci; ACT1 microsatellite, amylase (PyAmy) and direct amplification of length polymorphism markers, were applied (n = 163). Nevertheless, significant genetic differentiation was mainly observed owing to the PvAmy polymorphism (10 alleles, FST = 0.324) (Wanna et al., 2005). Accordingly, additional population genetic studies against large sample sizes are required for accurate estimation of genetic diversity and population differentiation patterns of P. merguiensis in Thai waters.
In Thailand, domestication of P. merguiensis has been carried out for production of high-quality pond-reared broodstock since 2000s (Putth Songsangjinda, personal communication). However, only a few parents of shrimp from Trang were recruited with intermittingly introduced of new brooders of the same origin to the breeding facilities. Limited genetic diversity of this stock after hatchery-propagated for 21 generations was recently reported by microsatellite analysis (average number of alleles per locus, NA = 4.857 and observed heterozygosity, Ho = 0.462; Praserlux et al., 2024).
Genetic improvement for increasing growth performance is required by the shrimp industry (Prasertlux et al., 2024). Identification of current genetic diversity and population structure of wild P. merguiensis in Thai waters is the crucial step for the re-establishment of a breeding program of P. merguiensis where its gene pools have been severely affected from overexploitation by conventional fisheries. The genetic information has not been monitored for approximately 2 decades since the last two studies (Hualkasin et al., 2003; Wanna et al., 2005).
Population genetic studies of wild P. merguiensis in Thai waters based on COI polymorphism are performed in the present study. Wild shrimp from various geographic locations were collected and polymorphism of COI was evaluated. Knowledge about genetic diversity and geographic differentiation of wild P. merguiensis is useful for management of its natural resources in Thai waters and for selection of appropriate founder populations of our breeding program of P. merguiensis.
Materials and Methods
Wild banana shrimp (P. merguiensis) were caught from geographically different locations including Nakhon Si Thammarat (n = 28), Chonburi (n = 30), Pattani (n = 24), Rayong (n = 28) and Trat (n = 24) located in the Gulf of Thailand (east coast). Shrimp were also collected from Trang (n = 33) and Krabi (n = 29) located in the Andaman Sea (west of peninsular Thailand; Table 1 and Fig. 1). Taxonomic identification of P. merguiensis was carried out following Pendrey et al. (1999). Pleopods were dissected out from each shrimp, placed in absolute ethanol, and transported back to the laboratory at National Center for Genetic Engineering and Biotechnology (BIOTEC). Specimens were kept at –80°C until needed.

Genomic DNA was extracted from a piece of pleopod of each shrimp using a GF-1 Tissue DNA Extraction Kit following the protocol recommended by the manufacturer (Vivantis Technologies, Selagor). The extracted DNA was visualized by agarose gel electrophoresis (1.0%) and concentrations were estimated using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA). The genomic DNA was stored at 4°C until further analysis.
The universal primers LCO1490 (5’-GGTCAACAAATCATAAAGATATTGG-3’) and HCO2198 (5’-TAAACTTCAGGGTGACCAAAAAATCA-3’) (Folmer et al., 1994) were used for the amplification of the partial COI gene segment (709 bp) of P. merguiensis. The amplification reaction was performed in a 25 μl reaction volume containing 1X reaction buffer, 2.0 mM MgCl2, 200 μM each dATP, dTTP, dCTP and dGTP, 0.25 μM each primer, 1 unit Taq DNA polymerase (New England Biolabs, Ipswich, MA, USA) and 50 ng genomic DNA. Polymerase chain reaction (PCR) was carried out using conditions described in Khamnamtong et al. (2009). The PCR product (5 μl) was size-fractionated through a 1% agarose gel to determine the successful amplification of the desired gene. The amplified COI of P. merguiensis was purified using a QIAquick PCR Purification Kit (Qiagen, Hilden, Germany) and direct sequenced for both directions by an automated DNA sequencer (ABI3730, Macrogen, Korea).
The generated COI sequences from 196 shrimp were searched against previously deposited sequences in GenBank using BlastN (Altschul et al., 1990). Nucleotide sequences of COI of P. merguiensis (GenBank accession no. PQ230498–PQ230693) were multiple-aligned by ClustalW (Thompson et al., 1994). Haplotypes were generated from identical sequences. Relationships between COI haplotypes were analyzed using PopART (Leigh & Bryant, 2015). Nucleotide sequence divergence between pairs of mitotypes was calculated. Haplotype diversity (Hd) and nucleotide diversity (π) within samples and pairwise nucleotide diversity and divergence between populations were calculated (Nei, 1987) using DnaSP (Rozas et al., 2017). The selective neutrality tests were evaluated by Tajima’s D (Tajima, 1989), and Fu’s Fs (Fu, 1997) statistics for each locality using DnaSP. For phylogenetic analysis, COI sequence of the black tiger shrimp Penaeus monodon (GenBank accession no. MF563563.1) was retrieved and used as an outgroup. The original data were bootstrapped 1,000 times. A neighbor-joining tree (Saitou & Nei, 1987) was constructed from nucleotide divergence between pairs of COI sequences using MEGA11 (Tamura et al., 2021). In addition, a NJ tree between wild populations of P. merguiensis was also constructed from interpopulation nucleotide divergence using Phylip (Felsenstein, 1993), and illustrated using FIGTREE 1.4.4 (Rambaut, 2018). Pairwise FST (Weir & Cockerham, 1984) were analyzed. Analysis of molecular variance (AMOVA; Excoffier & Lischer, 2010) were applied to test for statistically significant differences between hierarchical levels (between individuals within populations and between populations within groups) on different groupings including between Gulf of Thailand and Andaman Sea, among upper (Chonburi, Rayong and Trat) and lower (Nakhon Si Thammarat and Pattani) Gulf of Thailand and Andaman Sea (Trang and Krabi), and among phylogenetic groups (cluster I, Chonburi, Rayong, Trat and Pattani; cluster II, Nakhon Si Thammarat and cluster III, Trang and Krabi) using Arlequin 3.5 (Excoffier & Lischer, 2010). The number of female migrants between pairs of geographic samples per generation (Nefm) was calculated using Nefm = (1 – FST) / 2 FST (Hudson et al., 1992).
Results
The COI gene segment significantly matched the previously deposited COI sequence of P. merguiensis in GenBank (E-value = 0.0). Multiple-sequence alignments of the COI gene across overall samples generated 38 haplotypes (Table 2 and Fig 2). In total, 95 polymorphic sites were found. Only haplotypes 1 (H1, 65 individuals accounting for 33.16% of examined shrimp), H30 (23 individuals, 11.73% of examined shrimp) and H32 (47 individuals, 23.98% of examined shrimp) possessed by > 10% of examined samples. These haplotypes are regarded as major haplotypic groups connected with 35 minor haplotypes. Of these, 33 haplotypes were found in only one population (Table 2).
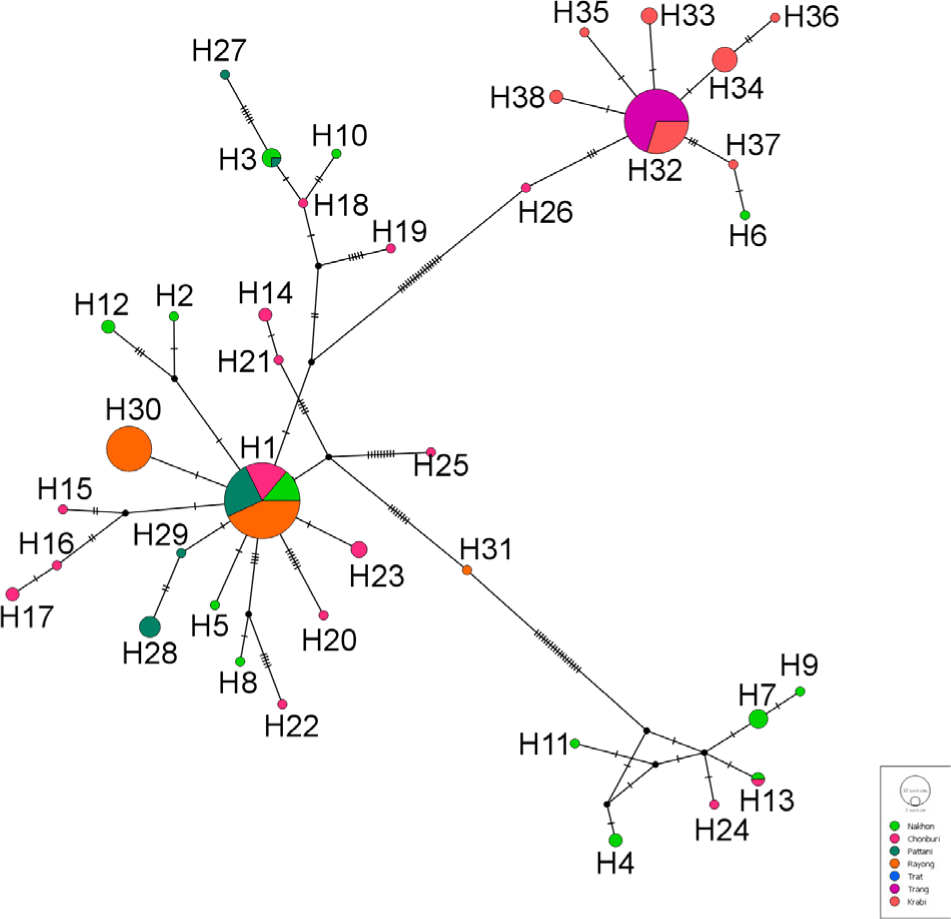
The H1 and H30 were majorly found in the east-coast samples (9/28, 12/30, 16/24 and 28/28 accounting for 32.14, 40.00, 66.67 and 100% in Nakhon Si Thammarat, Chonburi, Pattani and Rayong populations for H1 and 23/24 accounting for 95.83% in Trat for H30). These haplotypes were not distributed in Trang and Krabi (west). In contrast, the H32 were found in 100 (33/33) and 48.28% (14/29) of shrimp from Trang and Krabi but not in the east-coast samples.
High genetic diversity was observed in shrimp from Nakhon Si Thammarat (Hd = 0.876, π = 0.024) and Chonburi (Hd = 0.837, π = 0.013). A moderate haplotype diversity (Hd = 0.529) but a limited nucleotide diversity level (π = 0.003) was found in the Pattani population. Limited genetic diversity was found in Trat (Hd = 0.083, π = 0.001). Only one haplotype was found in Rayong (found only in the east-coast populations) and Trang (found only in the west-coast populations) reflecting a monomorphic haplotype and a lack of nucleotide diversity in these populations. The average haplotype and nucleotide diversity across overall populations were 0.818 ± 0.019 and 0.019 ± 0.001 (Table 3).
Microsatellite polymorphism at 7 loci and sample sizes (in brackets) are shown (Prasertlux et al., 2024).
Tajima’s D (Class I test; Tajima, 1989) which is analyzed based on the frequency spectrum of mutations and differences between estimators of the population mutation rate and Fu’s Fs (Class II test; Fu, 1997) which is calculated based on the haplotype distribution, were tested. Both Tajima’s D and Fu’s Fs were negative for overall populations of wild P. merguiensis. In these cases, results were not statistically significant (p > 0.05). Considering each population, Fu’s Fs estimates were not significant for all examined populations except Krabi (p < 0.05) while Tajima’s D was statistically significant in wild shrimp in Chonburi and Trat (p < 0.05) (Table 3).
Nucleotide divergence between pairs of wild geographic populations was 0.002–0.037 (Table 4). The lowest degree of interpopulation divergence was found between Trang and Krabi (0.001) located in the west-coastal region while larger nucleotide divergences were found between pairs of the remaining populations (0.002–0.037). The greatest degree of nucleotide divergence was found between Nakhon Si Thammarat and each of the Andaman populations (0.037) (Table 4).
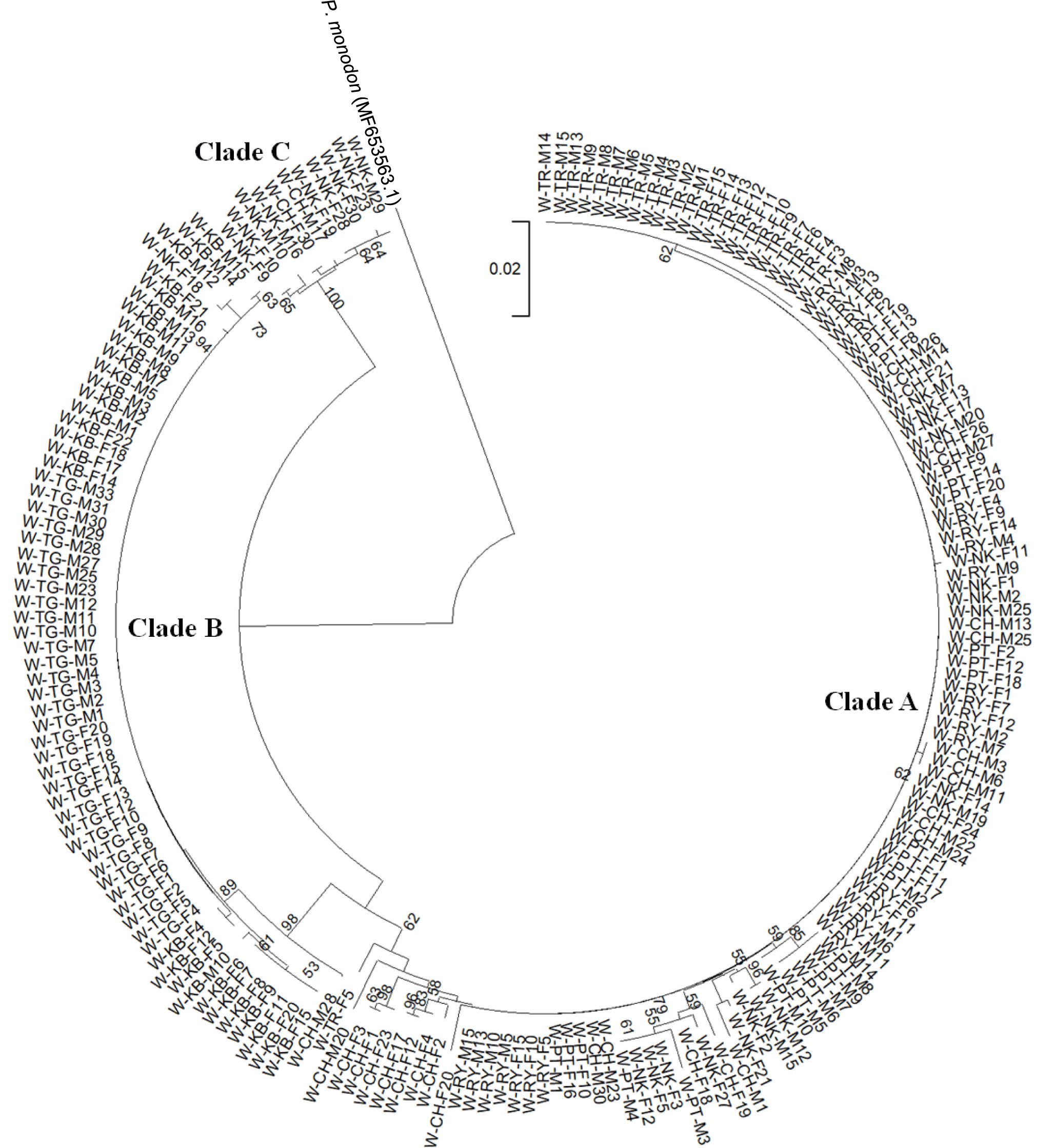
Phylogenetic analysis clearly revealed population subdivisions of wild P. merguiensis originated from different geographic locations. All shrimp from the west coast were allocated to phylogenetic clade B (n = 62 accounting for 32.14% of overall samples) while 117 and 15 shrimp originated from the east of peninsular Thailand (59.69 and 7.65% of overall samples) were phylogenetically allocated to clades A and C. Only two individuals, one from Nakhon SI Thammarat (W-NK-F18) and the other from Chonburi (W-CH-M28), were misplaced into clade B (Fig. 3).
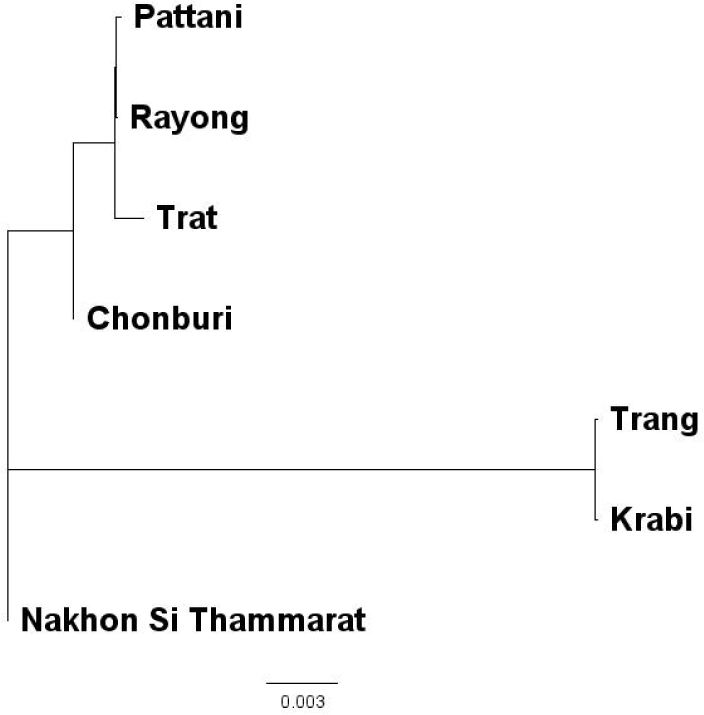
At the population level, a neighbor-joining tree allocated wild population of P. merguiensis in Thai waters to 3 different genetic clusters including Pattani, Rayong, Trat and Chonburi (cluster I), Nakhon Si Thammarat (cluster II), and Trang and Krabi (cluster III) (Fig. 4).
Pairwise FST estimates revealed significant population differentiation between all comparisons (p < 0.05) reflecting both between- and within coastal differentiation. A weak degree of population structure was observed between Nakhon Si Thammarat and Rayong (p < 0.05) while strong degrees of population subdivision were found between the remaining comparisons (p < 0.0001). The estimated female gene flow of wild population in this study was 0.040 (Chonburi and Krabi) –7.578 (Rayong and Trat) female migrants/generation. A moderate gene flow level was also observed between Rayong and Chonburi (7.500) located less than 100 km apart (Table 5).
When hierarchical structure was divided according to the coastal regions, AMOVA revealed highly significant differences between variance components among individuals within geographic populations (18.70%, FST = 0.8183, p < 0.0001) and among populations within coastal regions (5.15%, FSC = 0.2209, p < 0.0001) but not between coastal regions (76.78%, FCT = 0.7678, p > 0.025; Table 6). Likewise, genetic differentiation among groups was not found when upper and lower Gulf of Thailand were further divided and tested against the Andaman populations (69.17%, FCT = 0.76917, p > 0.0017). When hierarchical groups were divided following phylogenetic analysis (Fig. 4), significant differences between variance components were found among individuals within populations (22.33%, FST = 0.7767, p < 0.0001), among populations within groups (2.18%, FSC = 0.0890, p < 0.0001) and between phylogenetic groups (75.49%, FCT = 0.7549, p = 0.0059; Table 6).
AMOVA, Analysis of molecular variance; AS, Andaman Sea; GOT, Gulf of Thailand; df, degree of freedom; SS, sum of squares; VC, variance components; PV, percentage of variation, ns, not significant following Bonferroni’s adjustment (p < 0.025 for AS and GOT grouping and p < 0.0167 for AS, upper- and lower GOT grouping).
Discussion
The fundamental information on population genetics (genetic variability, population subdivisions and gene flow levels) is crucial for effective fishery management and sustainable breeding programs of P. merguiensis (Angel et al., 2022; Aziz et al., 2020; Chauhan & Rajiv, 2010; Hualkasin et al., 2003; Prasertlux et al., 2024; Wanna et al., 2005). The primary objective in this study is the application of population genetic information to assist the breeding program for improvement of growth performance of P. merguiensis.
Although high polymorphism of COI sequences was previously reported in P. merguiensis from different locations in Thai waters, genetic differentiation was only observed between shrimp from different coastal regions (i.e., Gulf of Thailand and the Andaman Sea) but not within the regions (Hualkasin et al., 2003). This probably due to limited sample sizes as only 26 individuals (n = 1–5 from each of seven geographic locations in peninsular Thailand) were included in the analysis.
In the present study, larger sample sizes were collected from seven different geographic locations (n = 196). Haplotype distribution frequencies of wild populations indicated region-specific haplotypes (H1 for shrimp from the east coast and H32 for shrimp from the west coast). Many singletons were observed in shrimp from each coastal region. High genetic polymorphism was observed in shrimp from the east of peninsular Thailand (Chonburi and Nakhon Si Thammarat) but low (or a lack of) either haplotype or nucleotide diversity of COI was found in Rayong (Hd = 0.000 and π = 0.000), Trat (Hd = 0.083 and π = 0.001) and Pattani (Hd = 0.529 and π = 0.003) in the east, and Trang (Hd = 0.000 and π = 0.000) and Krabi (Hd = 0.706 and π = 0.002) in the west of peninsular Thailand.
Using the same mtDNA region, a slightly lower genetic diversity (Hd = 0.54–0.62 and π = 0.001–0002) were found in green tiger prawn Penaeus semisulcatus from the Gulf of Suez and the Bitter Lakes, Egypt (Mohammed-Geba & Yousif, 2022) and Indo-Pacific region (Hd = 0.307–0.906 and π = 0.002–0027; Abdul Halim et al., 2021). In contrast, a greater diversity was found in Penaeus monodon (Hd = 0.716–0.927 and π = 0.029–0.085) originated from peninsular Thailand (Khamnamtong et al., 2009), Kuruma prawn Marsupenaeus japonicus (Hd = 0.993–1.000 and π = 0.029–0.045) from South China Sea and Taiwan (Shih et al., 2011) and Indian white shrimp Penaeus indicus (Hd = 0.837 and π = 0.07) from the Indian Ocean (Sajeela et al., 2019).
There is a pattern of increasing mitochondrial DNA diversity in wild P. merguiensis from Trat and Rayong in the upper Gulf of Thailand toward Chonburi and Nakhon Si Thammarat (lower Gulf of Thailand). Apparently, shrimp from Rayong and Trat have low diversity within populations (Hd = 0.000 & 0.083 and π = 0.000 & 0.001) but high genetic diversity was observed in Chonburi and Nakhon Si Thammarat populations (Hd = 0.876 & 0.837 and π = 0.024 & 0.013). Interestingly, the Pattani population possessed moderate levels of Hd (0.529) but low nucleotide diversity (0.003). This may imply that the interface between the east-west natural populations of P. merguiensis may be located near Pattani.
Based on both mtDNA (this study, Table 2), shrimp from Nakhon Si Thammarat (east) and Krabi (west) were chosen as founders for our breeding program. In addition, shrimp from Trang that exhibited a monomorphic western haplotype of mtDNA and those from Rayong that exhibited a monomorphic eastern haplotype of mtDNA were also included. Tajima’s D neutrality analysis was significant in wild populations from Chonburi (exhibiting the second highest genetic diversity level) and Trat (one of the populations exhibiting low diversity levels). These populations were not included as founders in our breeding program.
Interestingly, banana shrimp from Krabi displayed the west-coast types of mtDNA. Nevertheless, admixture analysis based on microsatellite data revealed that the Krabi population possessed the east-coast gene pool (Prasertlux et al., 2024). This suggests special characteristics of P. merguiensis from this geographic location. Accordingly, repeated sampling and population genetic analysis of the Krabi population should be performed in the future.
Haplotype distribution and the NJ tree based on nucleotide sequence divergence between pairs of COI sequences of wild P. merguiensis in this study revealed clear lineage separation between shrimp from different coastal regions (i.e., east and west coasts, clades A and B) of peninsular Thailand. Wild shrimp from the east coast exhibited a greater genetic diversity than the west coast (Andaman Sea) populations and an additional lineage (clade C) was differentiated from the main lineage for P. merguiensis. One possible explanation for the existence of different evolutionary lineages of mtDNA reflecting coastal differentiation of P. merguiensis in Thai waters is an ancient separation of mtDNA, which subsequently occurrence of several associated minor haplotypes. This may be resulted by limited gene flow levels of P. merguiensis caused by a major physical barrier, such as the main current in the Straits of Malacca, which moves from south to north throughout the year, and the major reversals in the monsoon-driven surface current systems of the Gulf of Thailand (Dale, 1956; Wanna et al., 2005).
A lack of (or extremely limited) mtDNA polymorphism in this study but high microsatellite polymorphism (Ho > 0.500; Prasertlux et al., 2024) of the same sample set is found in Thai P. merguiensis. This circumstance may result from the possible female founder and/or bottleneck effects (Canino et al., 2010). Subsequently, these geographic populations might have undergone a recent population expansion afterwards.
Similarly, population genetic studies of Penaeus indicus originating from 7 geographic locations covering Arabian Sea and Bay of Bengal were examined using 16S rDNA (n = 121). In total, 17 haplotypes with haplotype diversity = 0.0000–0.6667 and nucleotide diversity = 0.0000–0.0017 were found. AMOVA and pairwise FST analyses (except between Kanyakumari and Kakinada) indicated strong population subdivision of Penaeus indicus with overall FST value of 0.30684 (p < 0.05) (Angel et al., 2022).
In Australia, the biological stock structure of P. merguiensis may occur in the Northern Prawn Fishery (Larcombe et al., 2016) but their boundaries are unknown. The Western Australia and Queensland biological stocks are clearly separated but it is not known whether these are completely independent stocks or not (Tanimoto et al., 2006).
FST estimates revealed genetic differentiation between all pairwise comparisons of wild P. merguiensis. AMOVA further confirmed high genetic differentiation of P. merguiensis in Thai waters. Patterns of genetic population structure were concordant between both pairwise FST and AMOVA analyses when phylogenetic populations were regarded (Gulf of Thailand except Nakhon Si Thammarat, Andaman Sea and Nakhon Si Thammarat). Nucleotide divergence between examined populations and estimated female gene flow levels also support this circumstance.
Significant intraspecific population subdivisions of Thai P. merguiensis suggested that its gene pool is not panmictic but genetically differentiated to several local genetic stocks. Strong degrees of population differentiation at microgeographic scales (i.e., less than 100 km) in this species like that previously reported in blue swimming crab (Portunus pelagicus) in Thai waters analyzed by both nuclear DNA (AFLP) and mtDNA (COI) polymorphism (Khamnamtong et al., 2021; Klinbunga et al., 2007). In contrast, Penaeus monodon exhibited genetic differentiation between populations located in different coastal regions but not within the same region based on mtDNA polymorphism (Khamnamtong et al., 2009).
Levels of genetic diversity in wild P. merguiensis from various geographic locations revealed strong degrees of population differentiation. Apart from the major physical barrier (the main current in the Straits of Malacca and the surface current in Gulf of Thailand) described above, reproductive maturation of shrimp from different regions may promote degrees of genetic differentiation following seasonal reproductive variation in this species (Qureshi & Amanat, 2014). Moreover, rainfall is a major factor tightly associated with the emigration rates of P. merguiensis juveniles. Limited numbers of large-size shrimp migrate during low rainfall periods while mass migration of all shrimp sizes occurs during high rainfall periods (Staples, 1980). In the east of peninsular Thailand, the rainy season for the inner Gulf of Thailand (August–October for Rayong and Trat) and further south region (December–February for Nakhon Si Thammarat and Pattani) is in different periods (Hualkasin et al., 2003; Wanna et al., 2005). This circumstance should have promoted strong genetic differentiation in P. merguiensis.
The finding on local genetic differentiation between populations of Thai P. merguiensis were concordant between mtDNA (this study) and microsatellite (Prasertlux et al., 2024) analysis. For conservation and natural resource management, the banana shrimp in Thailand is heavily exploited by normal fisheries. Therefore, it is managed as a genetically homogeneous stock. In the present study, three phylogenetic groups of P. merguiensis were found. Haplotype distribution and FST statistics further illustrated that P. merguiensis originated from different provinces should be recognized as distinct genetic stocks. Therefore, they should be managed separately (Carvalho & Hauser, 1994; Wanna et al., 2005).
Overfishing can cause the extinction of local stocks and the possible loss of their genetic characteristics (Chauhan & Rajiv, 2010; Ovenden et al., 2015). Accordingly, Total Allowable Catch (Papa et al., 2020) should be implemented for P. merguiensis from upper Gulf of Thailand and Pattani, the Andaman Sea, and Nakhon Si Thammarat. In addition, sustainable catch limits (Papa et al., 2020) in Rayong and Trang (Hd = 0.000 and π = 0.000) should be considered. Levels of genetic diversity and population differentiation should be yearly monitored (Ovenden et al., 2015). Stock enhancement can be carried out to compensate overexploitation by using the local populations (Carvalho & Hauser, 1994; Chauhan & Rajiv, 2010) to maintain the major haplotypes (e.g., H1 for shrimp from the Gulf of Thailand, H30 for those from Trat and H33 from the Andaman Sea) and unique haplotypes found in each local stock. Consequent effects of stock enhancement should also be regularly evaluated. In addition, our on-going genetic improvement program has been developed. The domesticated stocks would significantly reduce the overexploitation of wild P. merguiensis at present.
For aquaculture purposes, the key success of domestication and selective breeding programs require P. merguiensis from different genetic stocks as founders (Aziz et al., 2020) to create subsequent generations with high genetic diversity. Unlike Penaeus monodon and L. vannamei, genetic improvement of P. merguiensis through selective breeding programs has not been implemented in Thailand. Based on high levels of between-population genetic differences, the information was applied for selection of appropriate sources of broodstock to establish a new breeding program (a collaborative project between Thai DOF and JICA-SATREPS, Japan) of P. merguiensis. An interpopulation crossing scheme between shrimp from Rayong, Nakhon Si Thammarat on the east-coast and Trang and Krabi from the west-coast region. For this objective, females from one population were artificially inseminated with male from a different population. Subsequently, 45, 47 and 51 families of the first, second and third generation (G1, G2 and G3) have been produced. The performance of commercially important traits (e.g., growth) between different families of P. merguiensis in our breeding program were evaluated. The heritability (h2) for growth following Animal linear mixed model (Gilmour et al., 2009) of the G1 population was 0.278 ± 0.060. The genetic gain after one generation of selection in G2 was 1.44 g and the performance test of G3 postlarvae was performed in two commercial farms. The information suggested the promising results for genetic improvement of P. merguiensis after three generations (P. Sae-Lim, unpublished data).
The reduced level of genetic diversity is a major concern for the sustainable breeding programs of aquaculture species (Aziz et al., 2020; Sajeela et al., 2019). Monitoring genetic diversity of domesticated stock is crucial for the success of our breeding program. A base population of P. merguiensis recruited from different genetic stocks represented the gene pool of P. merguiensis in Thai waters. In this study, higher genetic diversity of east coastal populations than west coastal populations of P. merguiensis were found. Analysis of COI polymorphism is useful to monitor the percentage contribution of shrimp exhibiting the east/west coast genotypes in our breeding program. The information on mtDNA polymorphism could be applied to assist our ongoing breeding program of P. merguiensis. This will elevate the selection efficiency for genetic improvement of this economically important species. In addition, dissemination of broodstock and postlarvae of genetically improved P. merguiensis to local farmers is being carried out. In the long term, these genetically improved stocks will contribute to the sustainability of the shrimp industry in Thailand.
