Background
Quantitative reverse transcription-PCR (RT-qPCR) has long been an established tool for the variety of gene expression assays owing to its cost effectiveness, sensitivity, reproducibility, and simplicity. This methodology has been commonly considered as the most reliable technique particularly for the quantitative validation of differentially expressed genes during a designed experimental treatment (Radonić et al. 2004; Udvardi et al. 2008). Several factors affecting RT-qPCR results include variables associated with qualitative and quantitative differences of input RNA templates as well as efficiency and specificity of reactions. For this reason, in most cases, the amplifications of internal controls (normalization controls) in parallel with the gene(s) of interest have been unavoidable to overcome the noise generated by the variables above (Kubista et al. 2006; Bustin et al. 2009).
Classically known housekeeping genes (HKGs) such as cytoskeletal β-actin (ACTB), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), and 18S rRNA genes have been commonly used as internal standard in RT-qPCR based on the belief that these HKGs do not change their expression levels irrespective of experimental stimulatory treatments. However, a considerable number of recent studies have also claimed that these HKGs may not always be stable depending on aquatic species; they could be modulated by certain stimulatory treatments with variable expression levels under different physiological conditions (Li & Shen 2013; Taylor et al. 2013; Yuan et al. 2014). Hence, careful validation of internal controls is now highly necessary prior to expression studies for target genes of interest. Different statistical software tools have become available with this purpose, and the potential utility of computational algorithms to select the best candidate reference gene has been broadly agreed upon in various gene expression studies (Vandesompele et al. 2002; Pfaffl et al. 2004; Silver et al. 2006).
Pacific abalone Haliotis discus hannai is one of the most commercially important mollusk species in Korean aquaculture domain. Because many abalone farms in Korea have used the sea-cage system in the coastal areas, waterborne toxicants existing in these areas have become serious concerns, affecting various physiologies of growing abalones (Wan et al. 2011; Park & Kim 2013). In particular, heavy metals are one of the critical pollutants in coastal areas, caused by various anthropogenic and/or industrial activities. Due to their high bioaccumulation capability, negative effects of heavy metals are often thought to be widespread at multilevels of the aquatic food chains (Türkmen et al. 2008; Silva-Aciares et al. 2011; Song et al. 2014). Aquatic invertebrates especially including the benthic mollusks having basically sedentary characteristics have been recognized as a useful platform not only to study the effects of metal toxicants on the physiology of marine animals but also to develop an effective early-warning bioindicator of metal pollution in the coastal environments. Heavy metals are also known as potent pro-oxidant effectors, consequently leading the potential deregulation of various cellular events in the exposed organisms (Amiard et al. 2006; Kim et al. 2007). Hence, comprehensive understanding on the relevant gene expression patterns in response to metal exposures would be a fundamental basis to gain a better insight into the protective mechanism of the exposed animals against metal toxicants (Kim et al. 2007; Wang & Rainbow 2005). Undoubtedly, the selection of suitable reference genes should be a prerequisite requirement for launching an expression analysis project regarding heavy metal exposures. However, despite its importance, the evaluation of stable reference genes from this abalone species in relation with heavy metal exposures has been little studied. Based on this need, the objective of this study was to validate different HKGs associated with heavy metal exposures as the initial step in environmental toxicogenomics of this abalone species.
Methods
Abalone specimens used in the present study were the stock that had been maintained in the Experimental Fish Culture Station of Pukyong National University (PKNU). During the pre-acclimation period (2 weeks), abalones have been maintained in rectangular (1 × 3 × 0.5 m = W × L × D) tanks equipped with protein skimmers Water temperature and dissolved oxygen throughout this period were adjusted to be 20 ± 1 °C and 8 ± 1 ppm, respectively. Abalones were fed with commercial diet for abalones three times a day on an ad libitum basis, and daily water exchange rate was 50 % using 1-μm-filtered seawater.
Three sets of heavy metal exposure treatments were performed. First, juvenile abalones (average body weight = 42.5 ± 3.4 g and shell length = 7.2 ± 0.5 cm) were exposed to cadmium (Cd), copper (Cu), or zinc (Zn). Heavy metal chemicals (analytical grade) were used (Sigma-Aldrich, St. Louis, MO, USA). Six individuals were assigned to one of three 50-L tanks containing 40 L seawater supplemented with each heavy metal. The dose strength of each heavy metal (Cd, Cu, or Zn) was 0.5 ppm as a nominal concentration. The non-exposed control group was also made identically except for the heavy metal. Two replicate tanks were prepared for each treatment group including the non-exposed control. Temperature (in-tank temperature) was adjusted to be ranged at 20 ± 1 °C with room air conditioner, and dissolved oxygen was maintained to be 8 ± 1 ppm with an air-diffuser-connected aerator. Exposure duration was 24 h. After 24 h, three individuals were randomly chosen from each replicate tank (i.e., six individuals per treatment) and tissues (gill and hepatopancreas) were surgically removed from each individual abalone. Tissues were immediately frozen on dry ice and stored at −80 °C until used for gene expression assay. Second, abalones (same sized as above) were exposed to different dose strengths (0, 0.1, 0.25, and 0.5 ppm) of Cu for 48 h. The preparation of two replicate tanks each containing six individuals and other tank conditions were the same with those used for the first exposure experiment. Three individuals were sampled from each replicate tank (n = 6 per treatment) in order to obtain the two tissue types (gill and hepatopancreas) from each individual. Third, abalones were exposed to Cd (0 or 0.25 ppm) for different durations of 24, 48, and 72 h. Twelve abalones were assigned into either one of four tanks (two replicate tanks for 0 and 0.25 ppm). Again, all other tank conditions were identically prepared with those mentioned above. During exposure treatments, no feed was provided, and the metal was renewed every 24 h with a daily water exchange rate of 20 %. At each time point, three individuals were randomly chosen from each replicate tank (six for Cd-exposed and six for non-exposed groups). Tissue sampling was carried out as described above. During the three exposure experiments, no mortality was observed.
Based on searches against both a local transcriptome next-generation sequencing (NGS) database constructed with abalone tissues (unpublished data) and the public NCBI GenBank database (http://www.ncbi.nlm.nih.gov), 12 candidate HKGs were selected. They included cytoskeletal β-actin (ACTB), β-tubulin (B-TU), elongation factor-1 alpha (EF1A), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), peptidyl-prolyl cis-trans isomerase B (PPIB), ubiquitin-conjugating enzyme E2 (UBE2), and six kinds of ribosomal proteins (RPL3, RPL4, RPL5, RPL7, RPL7A, and RPL8). Partial nucleotide sequences of the genes selected from the NGS database were confirmed by RT-PCR cloning followed by direct sequencing. Correct annotation for each gene from the NGS database was confirmed by BLAST homology search against NCBI GenBank. For each gene, multiple primer pairs were designed considering primer length (20–22 bp), G+C contents (40–60 %), melting temperature (55–65 °C), and amplicon size (100–200 bp). The PCR efficiency (E) for each gene was estimated using the known formula [E = 10(−1/k) − 1, with k = slop] (Kubista et al. 2006), based on the standard curve prepared with 4-log serial dilution points of abalone complementary DNA (cDNA) samples. LightCycler 480 Real-Time PCR System (Roche Life Science, Mannheim, Germany) was used for all the qPCR experiments in this study. Based on a series of preliminary evaluations, the optimal pair of primers was selected for each gene for further use in expression stability analysis. The summarized information on the genes and oligonucleotide primers used in the present study is provided in Table 1. The amplification of a single specific product using the best primer pair optimized for each gene was confirmed with end-point RT-PCR (Fig. 1).
aPCR efficiency
bDetermination coefficient

Total RNA from either gill or hepatopancreas was extracted using TriPure Reagent (Roche Applied Science) and further purified by using RNeasy Plus Mini Kit (Qiagen, Hilden, Germany) and RNase-Free DNase Set Kit (Qiagen) according to the manufacturers’ instructions. The quality and quantity of purified total RNA was checked with Libar S70 spectrophotometer (Biochrom Ltd., Cambridge, UK) at 230, 260, and 280 nm, in which only the RNA samples showing the absorbance values greater than 1.8 for both A260/280 and A260/230 ratios were considered for use. The integrity of selected RNA sample was validated with 1 % MOPS formaldehyde agarose gel electrophoresis. The absence of genomic DNA contamination in the purified RNA samples was confirmed by no amplification product (the intronic fragment of abalone actin gene) in the PCR reaction without RT reaction (data not shown). An aliquot (2 μg) of total RNA from each sample was reverse transcribed into cDNA using Omniscript RT Kit (Qiagen) according to the manufacturer’s recommendation. The RT product (cDNA) was diluted fourfold with DNase-free distilled water and 2 μL was subjected to PCR amplification. Thermal cycling in a reaction volume of 20 μL was performed with Light Cycler 480 System (Roche Applied Science) and SYBR Green I Master (Roche Applied Science) under the following conditions: initial denaturation step (at 95 °C for 10 min) followed by 40 cycles of 20 s at 95 °C, 20 s at 58 °C, and 20 s at 72 °C with an initial denaturation step at 94 °C for 2 min. For each sample, triplicate reactions were performed. The specificity of the amplification product was verified by melting curve analysis with the default setting in the thermal cycler. All the samples were assayed three times for technical replications.
Raw quantification cycle (Cq) data obtained were divided into two main datasets on the basis of tissue types because the transcriptional responses of certain target gene(s) to metal-stimulated treatment should be usually interpreted in a tissue-specific fashion. In the present study, we selected gill and hepatopancreas tissues, the two main organs with respect to the uptake and bioprocess of heavy metal ions in aquatic organisms. In order to evaluate the expression stability of candidate reference genes in either tissue type, the first dataset comprised of gill samples (n = 84 including biological replications), while the second dataset consisted of 84 hepatopancreas samples from non-exposed and metal-exposed groups. For each biological sample (i.e., individual cDNA sample) from either tissue types, technical replications of the qPCR assay were made in triplicates to determine the median value of each biological sample, and afterward, the control cycle threshold (Ct) values (i.e., biological replications) within an exposure treatment set were averaged in order to prevent the proportion of data from non-exposed control groups from being too high in the raw datasets.
To evaluate the expression stability of the HKGs tested, the RT-qPCR data was statistically analyzed using different types of software programs, geNorm (qBasePLUS ver. 3.0) (Vandesompele et al. 2002; Hellemans et al. 2007), NormFinder (ver. 0.953) (Andersen et al. 2004), BestKeeper (Pfaffl et al. 2004), and comparative delta Ct (ΔCT) method (Silver et al. 2006). For geNorm, the lower Cq value (i.e., the maximum expression level) from each gene was used as a calibrator and was arbitrarily set = 1. The geNorm algorithm calculates an expression stability value (M) for each gene and then compares the pairwise variation (V) of this gene with the others. The cutoff levels for M and V values were set to 1.5 and 0.15, respectively. Candidate genes with the lowest M value were considered to be the most stable under tested experimental conditions. The minimum number of reference genes for accurate normalization was also estimated by the V value in the geNorm program. For NormFinder, the same input file format as the geNorm was adopted; however, Cq values were converted into relative quantities after correcting the PCR efficiencies. For BestKeeper, untransformed Ct values and PCR efficiency (E) were used to determine the best suited standards and to combine them into an index by the coefficient of determination and the P value based on the geometric mean of the Cq values. The ΔCT method compares the relative expression of pairs of genes within each sample to confidently identify useful HKGs. Finally, comprehensive ranking to identify the most appropriate control gene was determined using the software RefFinder (http://fulxie.0fees.us/?type=reference&ckattempt=1), in which the four abovementioned software programs were integrated to compare the ranking of the tested candidate reference genes. Based on the rankings from each program, RefFinder assigns an appropriate weight to an individual gene and calculates the geometric mean of their weights for the overall final ranking.
Based on comprehensive rankings of the candidate reference genes, the suitability of selected reference genes was validated with the gene expression assay of metallothionein (MT) in response to heavy metal exposure treatment. The gill and hepatopancreas cDNA samples both from the Cd-exposed (treated at 0.25 ppm Cd for 24 h) and non-exposed control groups were subjected to qPCR quantification of MT transcripts with different normalization genes. A segment of MT cDNA (GenBank accession no. KT895222) was amplified by qPCR using a pair of primers (MT-FW: 5′-GGTACCGACTGCAAGTGTAA-3′ and MT-RV: 5′-TCATCGGAAGTCATGTGAGC-3′; amplicon size = 189 bp; E = 1.94) under the amplification conditions as described above for reference genes. The expression level of MT transcripts was normalized against the most stable reference gene (ranked to the top; R1), the two most stable reference genes (R1+R2), the second unstable reference gene (R11), or the least stable gene (ranked to the bottom; R12). Expression levels of MT transcripts in the gill and hepatopancreas tissues from the Cd-exposed abalones were presented as the fold difference to those from non-exposed control group using the formula 2−ΔΔCt (Schmittgen & Livak 2008) based on the normalization of MT transcript levels against the expression levels of its own reference genes (R1, R1+R2, R11, and R12). The effect of the reference gene selection was examined if differential levels of MT transcripts could be presented depending on the reference genes selected. Calculated levels of MT transcripts in gill and hepatopancreas based on each normalization regime were subjected to one-way ANOVA followed by Duncan’s multiple range tests in order to examine statistical significance at P = 0.05.
Results
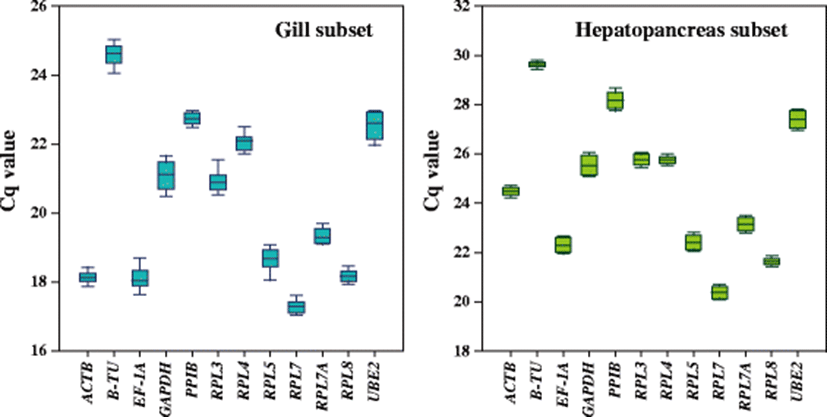
From technical and biological replicates, the efficiency of RT-qPCR calculated for each candidate reference gene was proven to range from 1.911 (RPL3) to 2.013 (RPL4) (Table 1). Also, every primer pair used in the present qPCR assays successfully revealed a single peak from the melting curve analysis (data not shown), indicating that the specific amplification for each candidate gene segment could be achieved. Of the 12 candidate reference genes tested, B-TU was the least expressed (the highest mean Cq values) gene in the gill subsets, whereas RPL7 was the most expressed gene in this tissue type. Considering the variation among samples in a given gill tissue, B-TU, EF1A, GAPDH, RPL3, RPL5, and UBE2 exhibited relatively greater variation in their expression levels than other candidate genes. In the subset of the hepatopancreas also, B-TU and RPL7 showed the least and highest expression levels, respectively, as similarly in the gill, although their absolute mean Cq values were different from those observed in the subset of the gill (Fig. 2).
Based on the evaluation with four different statistical approaches (geNorm, NormFinder, BestKeeper, and ΔCT method) followed by ranking decision with the RefFinder tool, all the four approaches did not recommend the same reference gene for the most suitable internal control gene (Table 2). According to the geNorm algorithm, the RPL8 was the most stable candidate gene in the gill, followed by RPL7 and ACTB. On the other hand, geNorm pointed B-TU and UBE2 as the two most unstable HKGs in the gill related with heavy metal exposures. NormFinder also recommended RPL7, followed by RPL8, for the most stable references while indicated B-TU as the most variable gene. According to BestKeeper, RPL7 and RPL8 turned out to be the most stable in the gill, whereas B-TU and UBE2 were pointed as the least stable candidates, although all the 12 genes tested showed SD values lower than 1. For the most stable candidate genes, according to the ΔCT method, the RPL7 gene was the first gene of the rank in the gill, while B-TU was the last gene. Collectively, based on the geometric mean-based ranking with RefFinder integrating the four statistical approaches, comprehensive ranking of each candidate gene was provided according to their stability, in which genes with lower ranking were considered to be most stably or invariantly expressed in the gill under heavy metal exposure conditions. As shown in Table 2, the recommended comprehensive ranking of the stability in the gill was determined as RPL7 > RPL8 > ACTB > RPL3 > PPIB > RPL7A > EF1A > RPL4 > GAPDH > RPL5 > UBE2 > B-TU. Overall, the results indicate RPL7 could be considered as the ideal reference gene in the gill regarding metal exposure experiment in this abalone species, while UBE2 and B-TU would be the worst or unstable references that are not recommended for the use in this case.
The stability ranking of candidate reference genes in the subset for the hepatopancreas was not exactly the same with that ordered in the gill, although the most stable reference gene finally recommended by the RefFinder program was the RPL7 in both tissue types (Table 3). With the geNorm program, the most stable candidates were RPL8 and ACTB while B-TU and GAPDH were the most variable references. NormFinder also pointed GAPDH as one of the least stable candidates whereas RPL7 and RPL3 were recommended as very stable references. On the other hand, BestKeeper showed slightly different patterns of rankings in the sense that B-TU, which was recognized as one of the least stable genes in other algorithms, was recommended as the most stable gene. This highest stability was followed by RPL8. However, for the least stable candidate reference genes, the GAPDH was called again by the BestKeeper approach. According to the ΔCT method, the most stable candidate was RPL7 while the most unstable gene was B-TU. Taken together, the comprehensive ranking determined by RefFinder for the subset in hepatopancreas was RPL7 > RPL3 > RPL8 > ACTB > RPL4 > EF1A > RPL5 > RPL7A > B-TU > UBE2 > PPIB > GAPDH.
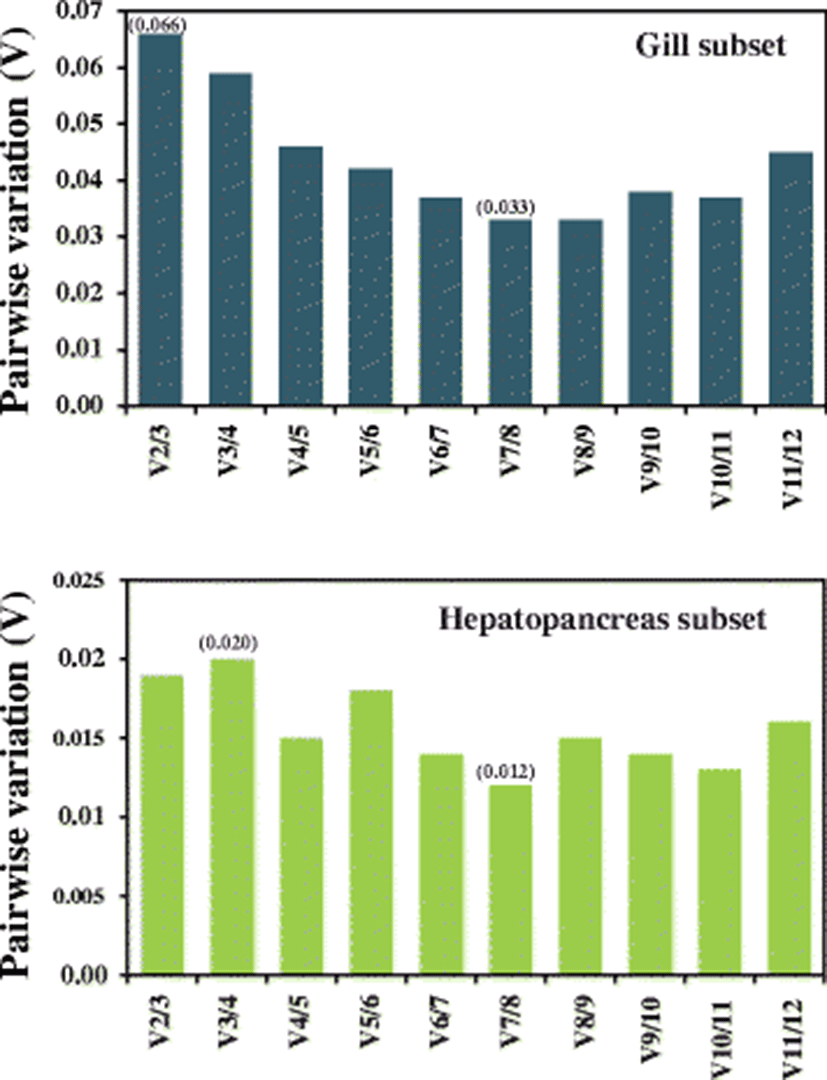
With the geNorm software program, the minimum number of reference genes for accurate normalization was determined based on the calculation of the pairwise variation Vn/Vn + 1 between two sequential normalization factors to consider the necessity of adding the next reference gene, in which large variation means that the newly added gene should have a significant effect on the normalization accuracy and hence be added for calculation. Considering the recommended cutoff value of 0.15, the V2/V3 value (0.066) observed in the gill criterion was much lower than the proposed cutoff value, suggesting that two reference genes are sufficient enough for the precise normalization of gene expression data in the gill samples regarding heavy metal exposures. Pairwise variation in the hepatopancreas subset also indicated that two reference genes could be very sufficient for accurate normalization in relation with the metal exposures in the present study, as evidenced by the V2/V3 value of 0.020 (cutoff value = 0.15) (Fig. 3).
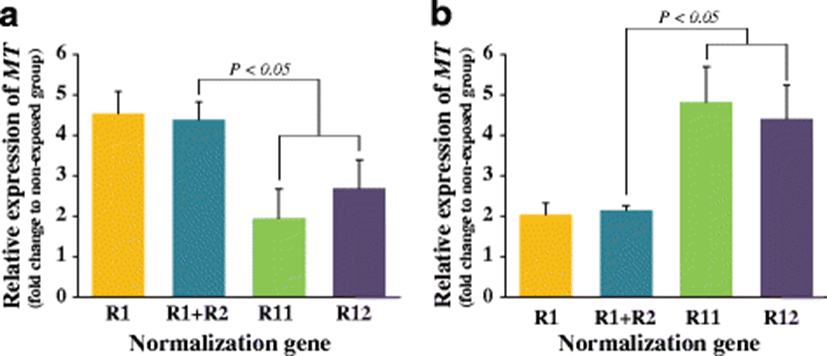
Stable or unstable reference genes showed different results in the interpretation of transcriptional responses of the MT gene to the Cd exposure (Fig. 4). In the gill, the most stable reference (R1; RPL7) and the combination of the two most stable references (R1+R2; RPL7+RPL8 based on the geNorm analysis) resulted in the similar amounts of MT transcripts induced during exposure (about 4.4-fold relative to non-exposed control group) (P > 0.05). However, when normalized to R11 (GAPDH) or R12 (B-TU), the calculated amounts of the MT transcripts in the gill of Cd-exposed abalones were only less than threefold relative to non-exposed controls (P < 0.05). In the hepatopancreas also, differential levels of the MT gene expression were visualized depending upon the reference genes selected. Similar with the finding from the gill tissue, the hepatic MT expression levels normalized to either R1 (RPL7) or R1+R2 (RPL7+RPL3) were similar with each other (about twofold relative to non-exposed control) (P > 0.05). However, the normalization regime using R11 (PPIB) or R12 (GAPDH) calculated the induced fold of MT transcripts in hepatopancreas to be more than fourfold as compared to the basal expression observed in the non-exposed control group (P < 0.05).
Discussion
Recently, the necessity to evaluate the stability of reference genes themselves prior to RT-qPCR assay for target genes of interest has been claimed by a number of studies to report that traditional reference transcripts may not always show constitutive expression as has often been assumed. It has already been widely agreed that no given single reference gene can be universally applied to all the experimental conditions and the utility of certain candidate reference genes could significantly vary among biotic and abiotic parameters including species, developmental stages, tissue types, and kinds of experimental treatments (Guénin et al. 2009; De Santis et al. 2011). For this reason, the case-specific choice of best reference gene for RT-qPCR may be unavoidable in many situations. In our case, we aimed to validate suitable reference genes from two subsets of tissue types (gill and hepatopancreas) in abalone regarding heavy metal exposures. Regarding gene expression studies of abalone tissues in response to heavy metal exposures, the gill is one of the most important biological barriers between internal and external environments of aquatic animals (Wang & Rainbow 2005; Hwang et al. 2011), and the hepatopancreas is the main organ responsible for deposits and metabolic detoxification of trace metals in aquatic invertebrates (Rainbow 2007). In this study, we performed two sets of heavy metal exposures (i.e., exposure to different heavy metals, Cu, Zn, Cd, and exposure to different doses of a given metal, Cu), and the raw RT-qPCR data were pooled into two subsets of tissue types (gill and hepatopancreas) in order to evaluate the expression stability of candidate reference genes in a tissue-specific fashion. The exposure conditions similar with those in the present study have already been proven to induce pro-oxidant stresses in abalone species belonging to Haliotidae (Kim et al. 2007).
For the reliable RT-qPCR, the high efficiency and specificity of PCR should be fulfilled. In our case, the calculated PCR efficiency (E) value for each candidate gene tested was at least higher than 90 %. Recommended efficiency values vary between 90 and 110 % (Schmittgen & Livak 2008; Doak & Zair 2012), and high E has been reported to be usually correlated with robust and precise interpretation of the gene expression (Bustin et al. 2009). The presence of a single peak for melting curve analysis with each PCR reaction also suggests that the specific amplification is carried out in the present conditions. Consequently, our RT-qPCR assay conditions comply well with typically known recommendations.
In the present study, we choose a total of 12 candidate reference genes including traditionally used genes (i.e., ACTB and GAPDH) but excluded the 18S rRNA, one of the most popular reference genes. The reason of exclusion was that 18S rRNA is not a poly(A+)-tailed RNA; therefore, the random priming for RT reaction is, at least in part, mandatory and the oligo-d(T) priming may not be applicable to this RNA type. In this case, due to its extremely high abundance, signal saturation obtained at very low Ct values might often be problematic unless template cDNA for the samples analyzed with 18S primers were diluted at least a hundredfold relative to all other candidate genes. Further, such a dilution procedure may cause a risk to introduce a random element of variability (Le et al. 2012). Alternatively, the 18S rRNA template could be prepared during an oligo-d(T)-primed RT reaction by adding an 18S rRNA-specific reverse primer at a very low final concentration (also called Co-RT method) (Zhu & Altmann 2005). However, in this case also, potential variability related with differential reverse transcription efficiencies between poly(A+)-tailed mRNA and abundant 18S rRNA should be taken into account. Although 18S rRNA has still been one of the versatile calibrators in many gene expression studies, many other studies also have argued that 18S rRNA should be considered as one of the not-recommended references in organisms belonging to a wide array of taxonomic positions (Taylor et al. 2013; Yuan et al. 2014; Shen et al. 2010). Twelve candidates selected in the present study are assigned to several categories of functional ontology such as cytoskeletal cell shape and mobility (ACTB and B-TU), energy metabolism (GAPDH), translation and protein synthesis (EF1A and various RPLs), and protein metabolism and degradation (UBE2). Such a selection regime might also be of importance for adopting software programs and interpreting outputs, since the algorithms used by each program differ from one another. For example, the geNorm usually requires a gene number ≥7 for a more reliable estimation and has the potential to detect co-regulated genes because its estimate is based on the pairwise comparison of the similarity of all the reference genes concerned (Vandesompele et al. 2002). On the other hand, the NormFinder calculates the variations within groups and between groups and it is less likely to detect co-regulation (Andersen et al. 2004). According to these, for more reliability, enough number of reference genes potentially belonging to different functional categories is recommended, and our selection regime is congruent with this requirement.
From expression stability tests, the two subsets of tissue types (gill and hepatopancreas) for abalones under either metal-exposed or non-exposed conditions shared a similar ranking pattern as judged by RefFinder in the sense that both subsets recommended equally the RPL7 as the most stable candidate gene while pointing the traditional candidate GAPDH as a very unstable (second most unstable and the first most unstable in the gill and hepatopancreas, respectively) reference gene. Our finding on the high stability of RPL7 reference genes is in accordance with previous reports with aquatic animals including oyster (Crassostrea gigas; Du et al. 2013), medaka (Oryzias latipes; Zhang & Hu 2007), and halibut (Hippoglossus hippoglossus; Øvergård et al. 2010), although the RPL7 gene has been little recommended in abalone species so far. Meanwhile, the other ribosomal protein gene RPL5 has been recommended as the most stable gene in different-sized groups of red abalone (Haliotis rufescens; Lόpez-Landavery et al. 2014) and also disk abalone (Haliotis discus discus) broodstock exposed to xenobiotics (Wan et al. 2011). In the present study, RPL5 showed a clear tendency of tissue dependency in its stability during metal exposures where it was ranked as one of the least stable candidates in the gill; in contrast, it was the third most stable gene in the hepatopancreas. It suggests the importance of validating reference genes according to not only the experimental conditions but also tissue types under evaluation. GAPDH, the unstable gene with substantial variations in this study, encodes a known glycolytic enzyme responsible for carbohydrate metabolism. Due to its classical housekeeping role, GAPDH mRNA expression has been commonly recognized as an invariant internal standard. However, from the last decades, substantial claims have arisen to report that GAPDH is no longer a classical HKG that is involved solely with carbohydrate metabolism. Instead, GAPDH protein has been now recognized as a multiplayer associated with diverse cellular pathways especially including stress response, apoptosis, and innate immunity (Cho et al. 2008; Lee et al. 2013). Further, from aquatic animals, the upstream regulatory regions (i.e., 5′-flanking regions) of the GAPDH gene have been reported to reveal binding motifs targeted by various transcription factors that have been known to be potentially involved in the stress response pathways and/or innate immunity, which is also congruent with the potential variability of GAPDH gene expression during experimental challenges or treatments (Lee et al. 2013). As previously known, heavy metals are a potent inducer for cellular oxidative stress often accompanied with inflammations and apoptotic changes in certain cell types; thus, metal exposure has the potential to modulate GAPDH genes in aquatic animals, as evidenced by the mud loach (Misgurnus mizolepis; Cypriniformes) (Cho et al. 2011).
On the other hand, ACTB was recommended as the third and fourth most stable gene in the gill and hepatopancreas, respectively. ACTB is one of the most common references used for gene expression experiments also in aquatic animals; however, the utility as an internal control has been reported to be largely controversial among studies. Notably, a recent study with different abalone species (red abalone H. rufescens) reported that ACTB was the least stable reference in the hepatopancreas (Lόpez-Landavery et al. 2014). The unstable feature of the ACTB gene was also reported in this abalone species during bacterial challenge (Qiu et al. 2013). The significantly different basal levels of the ACTB transcripts across different tissues in aquatic animals might also be, at least in part, supportive of our finding with this abalone species (Cho et al. 2011; Kim et al. 2008). Meanwhile, other two commonly known HKGs, B-TU and UBE2, did not receive high stability values in either the gill or hepatopancreas subset in this study, which is different from previous observations with other aquatic animals which report fairly good suitability of these HKGs as internal control standards under various experimental conditions (Fernandes et al. 2008; Zheng & Sun 2011). The importance of appropriate selection of the reference gene(s) for quantitative gene expression analysis regarding the exposure of heavy metal to abalone was clearly verified with the validation experiment using the MT gene conducted in this study. MT protein is a non-enzymatic multiplayer playing essential roles in metal homeostasis and detoxification, and due to its high inducibility at both the mRNA and protein levels, MT has been long recognized as a potential biomarker of heavy metal with aquatic organisms (Mao et al. 2012). In the present study, the amounts of heavy metal (Cd)-induced MT expression were differentially calculated depending on selected reference genes having different expression stabilities. Our data potentially suggest that the transcriptional response of MT could be either underestimated (as evidenced in the gill; see Fig. 4a) or overestimated (in the hepatopancreas; Fig. 4b) if “wrong” references were used, although the direction of transcriptional response (i.e., upregulation or downregulation) was not affected by the kinds of reference genes tested. Collectively, the overall findings from this study confirm not only that the reference genes chosen under a certain experimental treatment conditions may not be universally applicable to other conditions but also that expression stability of a given reference gene could not have consensus across tissue types.
Conclusions
Twelve potential candidate reference genes for RT-qPCR-based expression studies were evaluated in the gill and hepatopancreas tissues of abalone H. discus hannai under heavy metal exposure conditions. Based on multiple statistical algorithms, both gill and hepatopancreas subsets recommended ribosomal protein genes (particularly RPL7) as stable reference genes while traditionally known HKGs such as B-TU and/or GAPDH genes as inappropriate references. Our results also highlight the importance of selecting the suitable reference gene(s) for RT-qPCR studies, considering not only the experimental conditions but also tissue types under evaluation. Data from this study could be a good fundamental basis to guide future design of RT-qPCR studies with respect to metal regulation/detoxification and other related physiological aspects in this abalone species.
Abbreviations
ACTB, cytoskeletal β-actin; B-TU, β-tubulin; Cq, quantification cycle; Ct, cycle threshold; EF1A, elongation factor-1 alpha; GAPDH, glyceraldehyde-3-phosphate dehydrogenase; NGS, next-generation sequencing; PPIB, peptidyl-prolyl cis-trans isomerase B; RPL, ribosomal protein L isoform; RT-qPCR, quantitative reverse transcription-PCR; UBE2, ubiquitin-conjugating enzyme E2








