Introduction
Kudoa septempunctata has been known to be a parasite on fish muscles, digestive tract and kidneys, and has 6–7 polar cysts per spore, but does not affect fish physiology or survival (Matsukane et al., 2010). Some studies have reported that eating raw fish infected with K. septempunctata causes food poisoning in humans (Ahn et al., 2015; Jang et al., 2016). Kawai et al. (2012) analyzed 113 food poisoning patients in eight prefectures of Japan in 2010, and reported that the main cause was olive flounders infected with K. septempunctata. Since K. septempunctata was also detected in the feces of patients associated with consumption of sliced raw olive flounders, the Ministry of Health, Labor, and Welfare of Japan declared the K. septempunctata as a new causative agent of foodborne disease, and the associated foodborne illness was named Kudoa food poisoning (Harada et al., 2012; Jeon & Kim, 2016), which has a shorter incubation period of 2–20 hours compared to other food poisoning-causing pathogens (Matsukane et al., 2010). The genotypes of K. septempunctata are classified as three types: ST1, ST2, and ST3. Both ST1 and ST2 were found mainly in Japan, but ST3 is mostly known to be detected in Korea (Takeuchi et al., 2016). Some animal tests using Asian house shrew Suncus murinus found that 107 spores of K. septempunctata per 1 g, caused food poisoning symptoms such as vomiting and diarrhea (Kawai et al., 2012; Ohnishi et al., 2013). In the analysis on the residues of the food poisonings, more than 106 spores were found out of 1 g of olive flounders, and this concentration was set as the standard for food poisoning in Japan (Kawai et al., 2012).
So far some studies have been conducted to investigate the infection of K. septempunctata of olive flounders in Korea. Song et al. (2013) collected 143 olive flounders obtained from 26 fish farms in Jeju, and 4 samples out of 143 detected as K. septempunctata positive. Moreover, Song et al. (2014) collected 1,107 olive flounders from 89 fish farms in Jeju, Jeollanam-do, Gyeongnam, Gyeongbuk, and Gangwon-do in Korea, and 0.9% of the olive flounders in Jeju were only found to be K. septempunctata positive, but K. septempunctata were not detected in olive flounders in other regions. K. septempunctata infection was detected in olive flounders exported from Korea to Japan, so the Japanese government tightened quarantine inspection on olive flounders from Jeju in Korea, using 106 spores as the judgement standard (Jeon & Kim, 2016; Kim et al., 2015). As a result, the Ministry of Oceans and Fisheries of Korea has implemented Kudoa management measures for fish farms at the same level as Japan since April 2014. However, the actual concern is the food poisoning caused from eating sliced raw seafood such as sashimi or sushi with K. septempunctata infected. Not only in Korea but also in all of the world, the K. septempunctata standard for sashimi or sushi has not been established, which requires the management strategy for preventing the Kudoa food poisoning.
Within the authors’ knowledge, the previous studies have focused mainly on K. septempunctata infection of olive flounders from fish farms, and there has been no research on K. septempunctata-infected sashimi or sushi that people can eat right away. Therefore, this study is to investigate the contamination status of K. septempunctata in sashimi and sushi in Busan, Korea, where eating the raw seafood is traditional food culture, and provide basic data necessary for preventing and managing Kudoa food poisoning.
Materials and Methods
From January to December in 2020, 236 samples including 216 sashimi and 20 sushi made of olive flounders were collected from seafood markets and sashimi restaurants in Busan. The samples used in the study were purchased from the market by government employees of the Environmental Sanitation Department of 16 district offices in Busan. The samples were refrigerated and transported to the laboratory, and the experiments were conducted immediately.
According to the Korean Food Code, the K. septempunctata infection test in this paper is carried out as follows; (1) scratch the surface of the sashimi and sushi at least five places and obtain 1 g, (2) apply saline solution to it, (3) mash the bottom lightly with a flat surface, (4) centrifuge the solution through the mesh at 1,500×g at 4°C for 15 minutes, (5) dispose of the top part of the solution and add saline to the sediment and mix it to obtain a homogenized solution as a test solution. DNA was extracted according to the manual of the Fast DNA Spin Kit for Soil (Mpbio, Santa Ana, CA, USA) manufacturer. The extracted DNA was inserted into the PowerCheckTM Kudoa Real-time PCR Kit (Kogenbiotech, Seoul, Korea) and the Kudoa Screening Test was performed using the Real-time PCR (ABI 7500 Fast, Alameda, CA, USA), which was also followed by the manufacturer’s manual.
From the Real-time PCR results, if an amplification appears when the Ct value is less than 35, and if the quantitative value is 1.0 × 105 or higher, then we carried out a microscopic examination to check the spores of K. septempunctata as follows; (1) add 10 uL trypan blue solution to 10 uL test solution and mix it, (2) inject the obtained 20 uL solution into the hemocytometer, (3) examine the presence of Kudoa spores containing 6 to 7 cysts under a microscope (Carl zeiss, Oberkochen, Germany) and count them.
If the Real-time PCR test results show an amplification curve, then the nested PCR was carried out for detecting K. septempunctata using primers to detect 18S rDNA and 28S rDNA, according to Grabner et al. (2012) (Table 1). The target size was verified using Automatic Electrophoresis (QIAxcel advanced, Cologne, Germany), and we analyze the sequence obtained from the gene amplification PCR, and confirm whether it is equivalent to the standard strains of NCBI Blast (https://www.ncbi.nlm.nih.gov/Blast).
According to Takeuchi et al. (2015), we performed nested PCR for the genotype analysis of K. septempunctata under the conditions of 95°C for 3 minutes, 35 cycles of 95°C for 30 seconds, 55°C for 30 seconds, 68°C for 60 seconds, and 68°C for 5 minutes (Table 1). The gene sequence from PCR results is aligned by using the ClustalX BioEdit program, the genotype analysis was carried out by comparing the sequence with the standard strain and the sequence variation at a certain size (Table 2).
Results and Discussion
We conducted a Real-time PCR screening test on 236 samples of sashimi and sushi of the olive flounders. In order to perform the quantitative analysis, we made the calibration curve from the five standard reference materials for 1.0 × 107, 1.0 × 105, 1.0 × 104, 1.0 × 103 Kudoa DNA copy. As shown in Table 3, K. septempunctata was detected in 29 samples (13.4%) out of 216 sashimi, but not detected in sushi.
K. septempunctata was detected mainly in the summer, more specifically with 2 samples (0.8%) in June, 25 samples (10.6%) in July, and 2 samples (0.8%) in August. Using the Chi-square analysis, we confirmed statistically that the K. septempunctata detection rates vary significantly from month to month (p = 0.000). Kim et al. (2018) also found that Kudoa-related food poisoning cases in 2015 and 2016 were the most common in April, followed by October and November in Gyeonggi Province, and were the highest in May and followed by August in Korea as a whole. Conversely, Song et al. (2013) reported that K. septempunctata is detected throughout the year from January to December. Therefore, further research is needed on the monthly infection trend of K. septempunctata.
By analyzing the results of the Real-time PCR screening, we found that the number of copies of Kudoa rDNA in 29 samples varies from 5 × 103 to 2.6 × 108 per 1 g. By using the microscopy, we found K. septempunctata in 11 sashimi samples (Table 3). From the average value of the spores measured by four times in the hemocytometer, the number of the spores detected by K. septempunctata ranges from 1.0 × 104 to 9.0 × 105 spores per 1 g, all of which were less than 106 spores per 1 g of the standard for managing K. septempunctata in Japan.
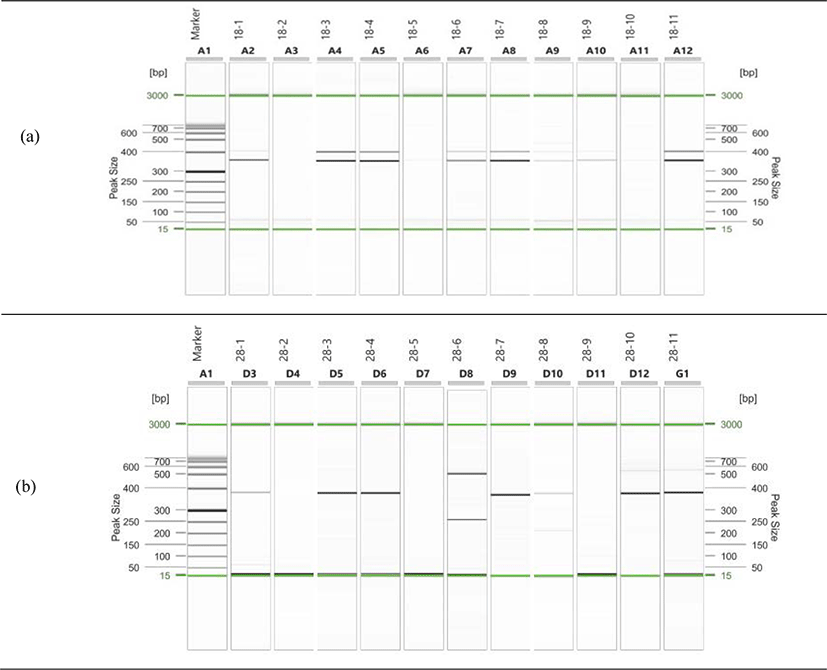
We conducted the PCR test on eleven samples with K. septempunctata spores observed by comparing with the 18S rDNA and 28S rDNA of K. septempunctata, and found that the 18S rDNA target size is 333 bp and the 28S rDNA target size is 356 bp, which is a specific part of the K. septempunctata (Fig. 1). Among the eleven samples, five (2.1%) samples were determined to be positive with K. septempunctata genes for nested PCR with both 18S rDNA and 28S rDNA. The distribution of olive flounders by original region is three from Jeju, one from Samcheonpo, and one from Wando (Table 4).
Song et al. (2013) surveyed 270 adult flounders and flounder fries from 26 fish farms in Jeju in 2012, and reported that 4 samples (2.8%) out of 143 adult flounders were detected with K. septempunctata genes. In addition, Song et al. (2014) collected 1,107 olive flounders from 89 fish farms in Jeju, Jeollanam-do, Gyeongnam, Gyeongbuk, and Gangwon-do, and reported that K. septempunctata genes were detected in 10 samples (3.1%) out of the 318 olive flounders in Jeju, but were not detected in other regions. Kim et al. (2015) surveyed 660 olive flounders in 11 fish seeding farms on the southwest coast in Korea between 2014 and 2015, and K. septempunctata genes were not detected. In contrast to previous papers, K. septempunctata were detected in the sashimi which made of olive flounders from Samcheonpo and Wando in addition to olive flounders from Jeju. In this study, a completely new result was found with the previous papers that K. septempunctata were detected only in olive flounders on Jeju. Therefore, further monitoring is needed for sashimi and sushi with olive flounders from other domestic regions.
In order to evaluate the gene consistency of K. septempunctata of the detected samples with the standard strains, we compared the gene sequence with 18S rDNA standard strains AB731754.1 and 28S rDNA standard strains AB731755.1. The five samples with K. septempunctata positive are identical from 98.4% to 100% consistent with the standard strains via NCBI Blast (https://www.ncbi.nlm.nih.gov/Blast) (Table 5).
The genotypes of K. septempunctata are ST1, ST2, and ST3, which are determined by combination of gene cox1 (cytochrome coxidase subunit 1) and gene rnl (large subunit rRNA). Gene cox1 is divided into cox1-1, cox1-2, and cox1-3 due to the differences of six nucleotides, while gene rnl is divided into rnl-1 and rnl-2 due to the differences of two nucleotides. By using Clustal W of the Bioedit program, we found that the gene sequences of K. septempunctata in the five samples are identical to ST3 with cox1-3 and rnl-2 (Fig. 2).

The genotypes of K. septempunctata vary depending on the country; ST1 and ST2 are mainly detected in Japan, while only ST3 is known to be detected in Korea (Takeuchi et al., 2015). In this study, the detected genotype of K. septempunctata is also ST3, which is consistent with the results of the existing studies; Kim et al. (2018) reported all the genotypes of K. septempunctata detected in food poisoning patients in Gyeonggi-do Province were ST3. However, Jang et al. (2016) could not find any connection with the foodborne diseases in animal tests by eating olive flounders with K. septempunctata of ST3. Since the infection path of K. septempunctata and its connection with the intermediate host has not been identified, the additional studies are needed to identify it. Eventually, further studies are needed on the connection between sliced raw olive flounders and Kudoa food poisoning (Ahn et al., 2015).
In the experiments, we first performed the Real-time PCR to screen the olive flounder samples, and then check the Kudoa spores with the microscopy, and finally determine whether they are positive by the PCR test. This study detected 29 samples to be K. septempunctata positive in the Real-time PCR, and the microscopy confirmed 11 samples among 29 samples, and the PCR tests finally confirmed 5 samples as positive which are detected from both 18S rDNA and 28S rDNA target of K. septempunctata. The number of detected samples decreased as they went through the steps, because the Real-time PCR test had unusual reactions or very little DNA in some samples (Grabner et al., 2012). In the future, the Loop-mediated isothermal amplification method (LAMP) or more advances in the Real-time PCR method is needed on preventing Kudoa food poisoning or tracking the infection path (Jeon & Kim, 2016; Kim et al., 2015; Song et al., 2014).
The previous researches only focused on the spread of K. septempunctata infections in olive flounders in fish farms, but no researches have been performed so far on the infection of K. septempunctata on sashimi and sushi in markets. In fact, the serious concern is that people have foodborne outbreaks from eating sashimi or sushi with K. septempunctata infected, so this study can be seen as the first step on preventing food poisoning. Now not only in Korea but also in the world, the detection criteria of K. septempunctata in sashimi and sushi have not been established yet, therefore we believe that the further research will be prioritized on the detection criteria for K. septempunctata in sashimi and sushi.
All the previous studies have detected K. septempunctata only in Jeju. However, this study found K. septempunctata in olive flounders sashimi from Samcheonpo and Wando other than Jeju. Within authors’ knowledge, this is the first study to detect K. septempunctata outside of Jeju. It is necessary to continuously monitor whether the detection region of K. septempunctata widen depending on climate change and environmental factors.








