Introduction
The gut of fish contains a very diverse and complex microbial community, which is known to affect the host’s intestinal development, immunity, and metabolism (Luan et al., 2023). When this balance is disrupted, fish are exposed to stress and can become diseased. It is generally known that the gut microbiota of fish is quite different from that of vertebrates such as mammals or birds (Kim et al., 2021; Ruiz-Rodríguez et al., 2020). In the human gut microbiota, Firmicutes and Bacteroidetes are dominant, whereas in fish, Proteobacteria is the dominant phylum (Kostic et al., 2013; Luan et al., 2023). However, our current understanding of the composition and role of the fish gut microbiota lags behind our knowledge of that of humans and other mammals (Luan et al., 2023). Many studies have reported that the gut microbiota of fish is involved in growth performance, digestive activity, physiological functions, and immune responses of fish (Hasan et al., 2018; Jang et al., 2023; Lee et al., 2024b). However, in most cases, the exact mechanisms are not well understood (Luan et al., 2023).
It is known that the gut microbiota of fish can be regulated by habitat temperature, environmental conditions, diet composition, and dietary supplements (Lee et al., 2023). Additionally, the diversity and composition of gut microbiota may vary depending on the growth stage of the fish (Lee et al., 2024a). Therefore, understanding the relationship between the fish gut microbiota and health, homeostasis, immune response, and fish dysbiosis opens up new possibilities for modulating the composition of the gut microbiota in farmed fish to produce healthy fish. However, this requires more information on the composition and diversity of the gut microbiota of fish, and understanding the relationship between fish and the gut microbiota can only be achieved when sufficient gut microbiota analysis results are available.
Olive flounder (Paralichthys olivaceus) is the most widely farmed fish species in Korea, accounting for approximately half of the total farmed fish production each year (Lee et al., 2024c). Olive flounder farms are concentrated in Jeju and Jeollanam-do, but are also found in Gyeongsangbuk-do and Gyeongsangnam-do. Regional differences in the environment and farming practices result in variations in the characteristics of farmed olive flounder. The Jeju region offers relatively favorable conditions for olive flounder farming, as it has access to underground seawater that can maintain a temperature of approximately 16°C–18°C, suitable for year-round cultivation. In contrast, the average water temperature in Jeollanam-do is about 1°C–2°C lower than that in Jeju, and during winter, it drops below the optimal temperature for olive flounder farming, requiring additional heating. In Gyeongsangbuk-do, the growth rate of olive flounder is relatively slow due to the large temperature fluctuations along the east coast (Baek et al., 2024). These regional environmental differences may affect the composition of the gut microbiota in olive flounder, potentially complicating the interpretation of associations between gut microbiota and fish health. Therefore, it is necessary to better understand these differences and accumulate comparative data on the gut microbiota of olive flounder across different aquaculture regions.
Given these regional environmental variations, this study aimed to investigate how aquaculture location influences the gut microbiota of similarly sized olive flounders. Specifically, alpha and beta diversity analyses were performed, along with an evaluation of the relative abundance of gut microorganisms across the regions.
Materials and Methods
Ten fish weighing approximately 100 g were collected from each of four locations where olive flounder was farmed (Fig. 1). The flounders were transferred to the laboratory, anesthetized with 2-phenoxyethanol (Sigma-Aldrich, St. Louis, MO, USA), weighed, and their entire intestines were aseptically collected immediately. Total DNA from intestinal microorganisms was extracted using the ZymoBIOMICS DNA Miniprep Kit (Zymo Research, Irvine, CA, USA) according to previously described method (Lee et al., 2024a).

Sequencing for microbiome analysis was performed using the Illumina MiSeq system (Illumina, San Diego, CA, USA) (NFEC-2025-03-305081), following the 16S Metagenomic Library Preparation Guide (Illumina et al., 2013). Briefly, the V3–V4 hypervariable regions of the 16S rRNA gene were amplified for amplicon sequencing. PCR amplification was conducted using primers with adapter overhang nucleotide sequences provided by Illumina. Subsequently, library quantification, normalization, and pooling were carried out according to the manufacturer’s instructions. Finally, sequencing was performed using the MiSeq sample loading protocol.
Data analysis was performed using Quantitative Insights Into Microbial Ecology 2 (QIIME2) v2023.5 (https://qiime2.org/) (Bolyen et al., 2019). Low-quality score regions (< 25) were trimmed using the Divisive Amplicon Denoising Algorithm 2 (DADA2) pipeline (https://github.com/benjjneb/dada2/), and subsequent merging, noise filtering, and chimera removal were conducted. Each read was clustered into amplicon sequence variants (ASVs) (Callahan et al., 2016) and taxonomic classification was assigned using the classify-sklearn method with the SILVA database v138 (https://www.arb-silva.de/). The analysis results were visualized using QIIME2 View (https://view.qiime2.org/) and MicrobiomeAnalyst (https://www.microbiomeanalyst.ca/) (Chong et al., 2020).
Results and Discussion
To compare the gut microbiota of olive flounder farmed in different regions of Korea, four representative aquaculture regions were selected based on their geographical characteristics. The selected regions include Gyeongsangbuk-do on the East Sea coast, Gyeongsangnam-do on the South Sea coast, Jeollanam-do on the Southwest Sea, and Jeju Island. The geographical coordinates (latitude and longitude) of each region are presented in Fig. 1.
In Korea, aquaculture is primarily concentrated in the West Sea, South Sea, and Jeju Island. For flatfish species, including olive flounder, the highest concentration of farms is located in Jeju Island (49.4%), followed by Jeollanam-do (30.0%) and Gyeongsangnam-do (7.6%) (Statistics Korea, 2024). In the Jeju region, 21,000 tons of flounder were produced in 2023, accounting for 52.5% of total production, while 17,000 tons (42.5%) and 2,000 tons (5.0%) were produced in Jeollanam-do and Gyeongsangnam-do, respectively (Statistics Korea, 2024). Although aquaculture is less developed along the East Coast compared to the West and South Seas, recent efforts led by the National Institute of Fisheries Science have contributed to the expansion of aquaculture technologies, resulting in the production of some aquaculture species in the region. Although production varies across regions, all four regions selected in this study currently operate flounder farms. These regions are expected to exhibit environmental differences due to their distinct geographic locations, particularly in terms of latitude and longitude. Therefore, in this study, these regions were selected as suitable sites for comparative analysis of the gut microbiota of flounder according to aquaculture region.
Table 1 shows the alpha diversity comparisons among groups based on three diversity indices. Group A exhibited the highest observed feature values. Statistical analysis revealed significant differences (p < 0.05) between Group A and Groups B and C. Among all groups, Group C showed the lowest observed feature values. For the Shannon index, significantly higher values were observed in Groups B and D compared to Group C (p < 0.05). Similarly, the Simpson index showed significantly higher values in Groups B and D than in Group C (p < 0.05). Fig. 2 presents the beta diversity analysis of olive flounder gut microbiota based on the Jaccard distance. Both 2D and 3D principal coordinate analysis (PCoA) plots showed clear separation among the groups. Samples within each group tended to cluster closely, with this pattern being particularly distinct in Groups A and D.
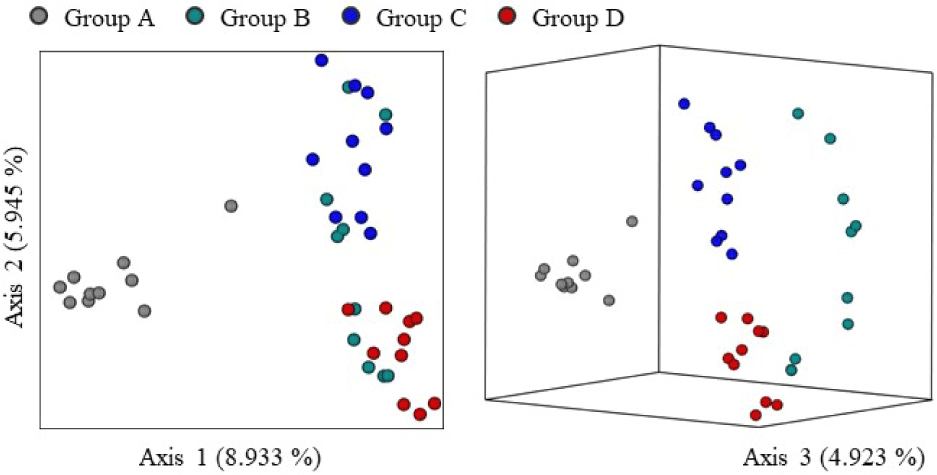
Our previous study reported that gut microbiota diversity varied depending on the growth stage of olive flounder. The results indicated that the gut microbiome of the subadult group (400–800 g) was more diverse and complex than that of the juvenile group (–100 g) (Lee et al., 2024a). Although this study compared the gut microbial communities of similarly sized olive flounders, differences in diversity were observed across aquaculture regions. Numerous previous studies have demonstrated that the composition of intestinal microbiota is strongly influenced by the host’s surrounding environment (Jang et al., 2021; Qiao et al., 2019). The nutritional status of flounders raised in different farms may vary depending on the composition and amount of feed provided. In addition, the temperature and microbial composition of seawater used for aquaculture may also differ by region. These factors may influence not only host health but also gut microbial diversity. However, the health status of olive flounder cannot be evaluated solely based on gut microbiota diversity. To establish a clearer relationship between gut microbiota composition and host health, further studies are needed to elucidate how specific microbial increases or decreases impact the host.
The results of relative microbial abundance at the phylum level, based on the SILVA database, are presented in Fig. 3. Although some groups exhibited similar microbial compositions, others showed notable differences in phylum distribution. The most abundant phylum in Groups B, C, and D was Proteobacteria. However, in Group A, Proteobacteria was the dominant phylum only in some individuals (A2, A4, A6, A8, A9, and A10), whereas Firmicutes was most abundant in the remaining individuals (A1, A3, A5, and A7). These within-group variations were most pronounced in Group A, but were also observed in Group B (B1 and B2) and Group C (C1). Additionally, Firmicutes was present in both Groups A and B, although at different relative abundances. Some individuals in Group C (C6 and C7) exhibited a relatively high abundance of Fusobacteriota.
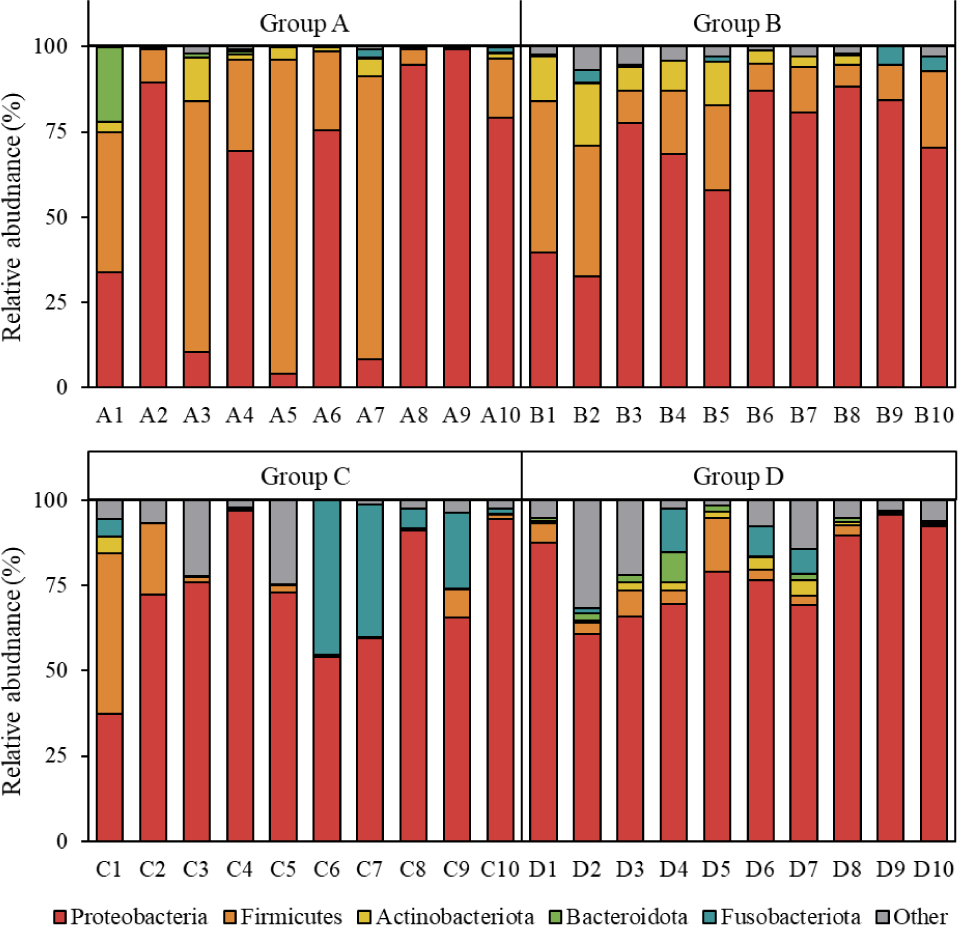
In our previous study, we reported that Proteobacteria was the most abundant phylum in the gut microbiota of juvenile olive flounder weighing less than 100 g (Jang et al., 2020; Lee et al., 2024a). Furthermore, as olive flounder grew, the relative abundance of Proteobacteria decreased, while that of Firmicutes increased (Lee et al., 2024a). Consistent with previous findings, most individuals in this study exhibited Proteobacteria as the dominant phylum, except for several individuals in Group A. In particular, some individuals in Group A, in which Proteobacteria was not dominant, showed the highest abundance of Firmicutes—known to increase with host growth. Therefore, at the phylum level, Proteobacteria was the most abundant across aquaculture regions, but the dominance of Firmicutes may vary depending on individual growth characteristics. However, this trend may differ in juvenile olive flounder fed with microbial additives such as probiotics.
The relative abundance of bacterial genera is shown in Fig. 4. In Group A, several individuals (A1, A2, A4, A6, A8, A9, and A10) showed a high relative abundance of the genus Vibrio. Additionally, in Group C, the genus Photobacterium was relatively abundant in most individuals, except for one (C3).
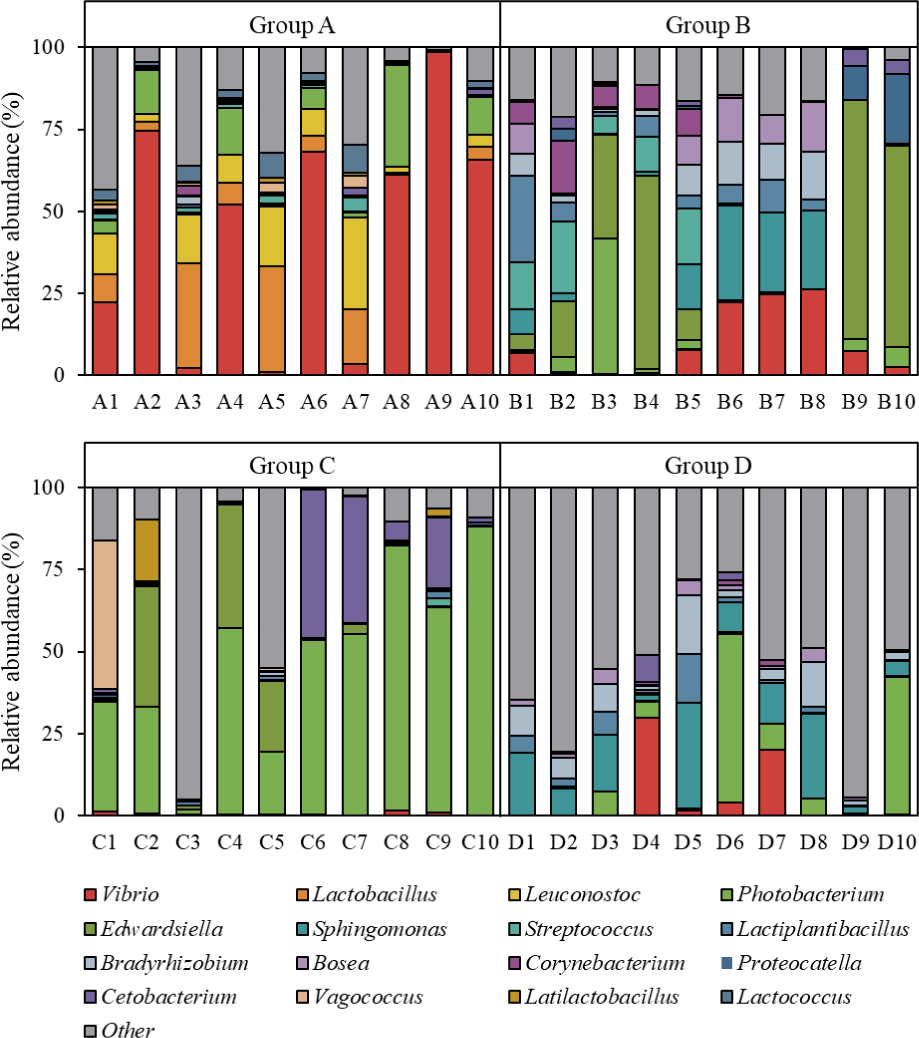
Key members of the fish gut microbiota are known to include Vibrio, Photobacterium, and Clostridium (Egerton et al., 2018). Similarly, Vibrio and Photobacterium have been reported as dominant genera in the gut microbiota of olive flounder (Lee et al., 2024a). In this study, these genera were abundantly present in certain groups (A and C), whereas Groups B and D did not exhibit high abundances of these taxa. Such differences are likely influenced by factors such as diet, feed additives, water quality, and water temperature. Therefore, further studies are needed to identify the factors affecting gut microbiota composition at the genus level to enable a more comprehensive interpretation.
The dominant genera in each group were identified using linear discriminant analysis (LDA) effect size (LEfSe) analysis. The genera with the highest LDA scores were Vibrio, Edwardsiella, Photobacterium, and Sphingomonas in Groups A, B, C, and D, respectively (Fig. 5).
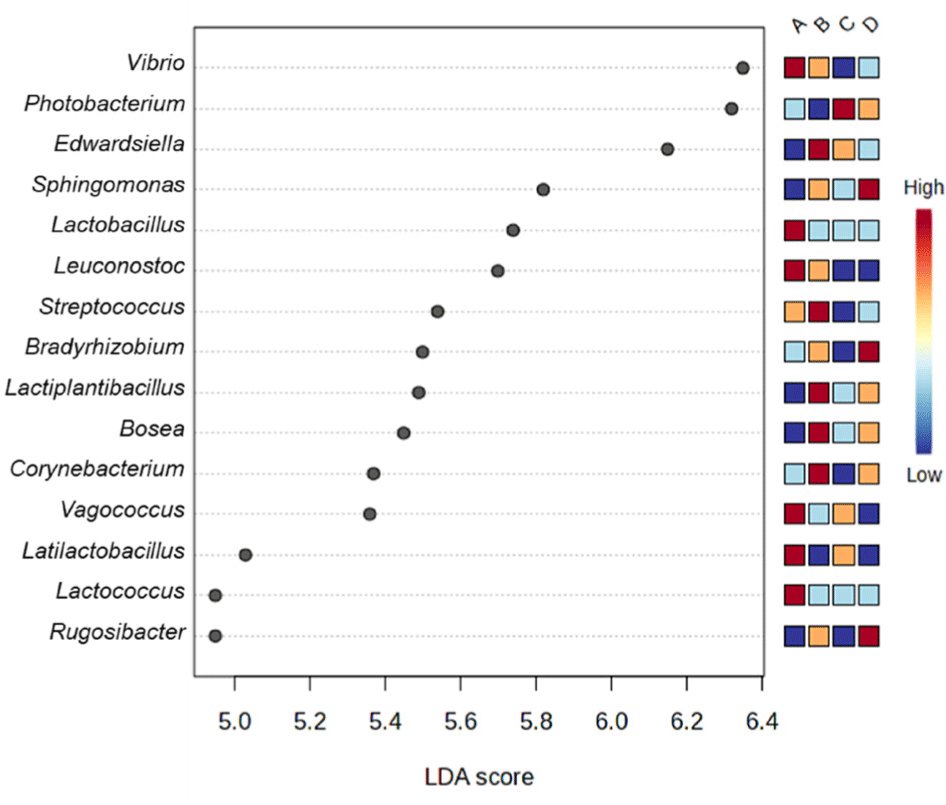
These results suggest that the composition of the gut microbiota in olive flounder varies depending on the aquaculture region. This further indicates that environmental factors associated with the farming environment may have a greater influence on gut microbiota composition than species specificity or the growth stage of the host.
Conclusion
This study confirmed that the composition of the gut microbiota in olive flounder at similar growth stages can vary depending on the aquaculture region. These findings suggest that gut microbiota composition is more strongly influenced by aquaculture environment and methodological factors than by the growth stage alone. Furthermore, the composition of the gut microbiota in farmed olive flounder may be modulated by adjusting factors such as feed type and the use of dietary additives. However, since this study focused only on flounder weighing approximately 100 g, further research is needed to investigate how environmental factors affect intestinal microbiota composition throughout the entire life cycle of olive flounder. A deeper understanding of these relationships will enable more precise control of intestinal microbiota composition, contributing to healthier and more sustainable flounder aquaculture.








