Introduction
The Indian seafood industry demonstrated robust growth in fiscal year 2022–2023, exporting 1.73 million metric tons of fish and fisheries products valued at $8.09 million, with squid contributing a notable 4.83% by quantity (PIB, 2023a). As the world’s third-largest producer, India accounts for 8% of global fish and fishery product output (PIB, 2023b). However, this success generates an equivalent amount of waste biomass from seafood processing plants (Olsen et al., 2014). This cephalopod-rich waste harbors significant potential due to its biologically active materials like chromatophore collagen, and eumelanin, (Nam et al., 2008). Both chromatophore and collagen have garnered substantial research interest for commercial exploitation, owing to their diverse properties (Sköld et al., 2016).
Recovered from cephalopods, collagen and chromatophore offer remarkable applications in the food, pharmaceutical, textile, and photographic industries, thanks to their unique properties such as ultraviolet ray absorption, antimicrobial activity, and protection against oxidative stress (Lewis et al., 2022). Chromatophore, a pigment sac containing ommochromes, typically requires a repeated recovery process involving homogenization with ultrasound, enzymatic hydrolysis, and acidified methanol (HCl-MeOH) extraction (DiBona et al., 2016). The efficacy of this extraction depends on factors like time, pH, temperature, and sodium chloride (NaCl) concentration (Oancea et al., 2012). Optimizing extraction methods, such as enzymatic and ultrasound treatments, is crucial for unlocking the full value of these marine byproducts. For instance, combining these techniques efficiently extracts highly antioxidant collagen hydrolysates from Alaska pollock skin, yielding fractions with biotechnological potential (Lee et al., 2022). Similarly, ommochrome pigments, specifically xanthommatin, is optimally extracted from jumbo squid skin through precise ultrasonication and temperature control, resulting in compounds with antibacterial and antimutagenic properties (Chan-Higuera et al., 2019). This enzyme-assisted approach not only maximizes yield but also minimizes environmental impact, representing a sustainable method for valorizing fishery byproducts (Feng et al., 2013). This highlights how targeted extraction of both collagen and ommochromes transforms marine waste into valuable resources for industrial application (Chan et al., 2019; Chen et al., 2023; Esparza-Espinoza et al., 2022).
The extraction and utilization of these valuable compounds from cephalopod processing waste not only demonstrate environmental responsibility but also prove economically viable. Current reports often employ multistage, repetitive steps for single-product recovery, such as ommochrome purification, which can be complicated by the presence of other abundant compounds like collagen, thereby neglecting the broader potential of these resources (Lewis et al., 2022). The strategic recovery of both ommochrome and collagen from seafood waste biomass offers multifaceted advantages across environmental, economic, and health and wellness dimensions. Our present objective is to develop a comprehensive two-stage purification process for both collagen and ommochromes extracted from Indian squid skin, involving the sequential separation of epidermis, collagen, and ommochromes.
To enhance the efficiency of these procedures, our analysis focuses on optimizing the sequential extraction of collagen and ommochromes from the chromatophore tissues of the Indian squid, Uroteuthis duvauceli. We employed traditional one-factor-at-a-time (OFAT) studies to identify significant factors affecting purification and obtain initial optimization conditions (Chauhan et al., 2021). For a more comprehensive understanding and to identify optimal conditions, we utilized response surface methodology (RSM), which is a statistical and mathematical approach considering multiple factors simultaneously (Reungoat et al., 2021). Furthermore, to enhance the reliability of our models and uncover complex, non-linear relationships, we adopted machine learning (ML) models such as artificial neural networks (ANN), Random Forest, and k-nearest neighbors (kNN) algorithm (Noi & Kappas, 2017). This investigation is the first of its kind to optimize the conditions for recovering both collagen and ommochromes from squid skin using a statistical approach, filling a significant gap in the existing knowledge base. We developed a second-order polynomial model for the recovery process and applied a similar methodology for ommochrome recovery. The purified collagen and nanostructured pigments from Indian squid pigment granules hold promising potential applications for both human well-being and the environment. Unlike conventional single-product, multi-stage recoveries, our novel two-stage, multi-product process from a single waste source uniquely enables the commercial recovery of valuable compounds like collagen and ommochrome, thereby enhancing industry income, reducing waste disposal costs, fostering ancillary industries, generating employment, improving fishermen’s socio-economic conditions, and contributing significantly to sustainable marine byproduct valorization.
Materials and Methods
All chemicals, enzymes, and reagents acquired for this study are of analytical grade and sourced exclusively from Merck Limited in Mumbai, India. Solutions and reagents were meticulously prepared in strict accordance with the current specifications outlined by the American Chemical Society (ACS) (1999).
Buffer solutions with pH values ranging from 1 to 9 were meticulously prepared for experimental use (Lambert & Muir, 1973; Robinson & Stokes, 2012). The solution at pH 1 was created by blending 50 mL of 0.2 M KCl with 134 mL of 0.2 M HCl. For pH 2, a mixture of 50 mL of 0.2 M KCl and 13 mL of 0.2 M HCl was employed. Achieving pH 3 involved combining 100 mL of 0.1 M potassium hydrogen phthalate with 44.6 mL of 0.1 M HCl. Buffering at pH 4 necessitated the combination of 100 mL of 0.1 M potassium hydrogen phthalate with 0.2 mL of 0.1 M HCl. To attain pH 5, a blend of 100 mL of 0.1 M potassium hydrogen phthalate and 45.2 mL of 0.1 M NaOH was prepared. The solution at pH 6 was crafted by mixing 100 mL of 0.1 M KH2PO4 with 11.2 mL of 0.1 M NaOH. Buffering at pH 7 involved combining 100 mL of 0.1 M KH2PO4 with 58.2 mL of 0.1 M NaOH. For pH 8, 100 mL of 0.1 M tris (hydroxymethyl) aminomethane and 58.4 mL of 0.1 M HCl were added. Finally, pH 9 was achieved by blending 100 mL of 0.1 M tris aminomethane with 11.4 mL of 0.1 M HCl.
A NaCl solution was prepared by weighing 0, 2, 4, 6, 8, and 10 g of NaCl and adjusting the volume to 100 mL in a volumetric flask, resulting in 0%, 2%, 4%, 6%, 8%, and 10% NaCl solutions.
Freshly prepared HCl-MeOH is obtained by carefully adding 0.010, 0.075, 0.550, 4.150, or 30.750 mL of concentrated hydrochloric acid (36.5%–38% HCl, ACS Reagent Grade) to 1,000 mL of cold methanol (high-performance liquid chromatography [HPLC] Grade MeOH), resulting in a methanol of pH of 9, 7, 5, 3, 1. Methanal without any added HCl is used as bank.
The Indian squid was harvested from the waters adjacent to the Undivided Dakshina Kannada District at coordinates latitude 13.21505 and longitude 75.05525. The squid exhibited a length range of 14–19 cm, with sample weights varying between 86 and 98 g. Following capture, the squid specimens were promptly immersed in chilled seawater at +4°C and transported to the department of biotechnology at P. A. College of Engineering within 45 minutes, utilizing an insulated container. Subsequently, they were stored at –20°C, in the event of any delays. The samples were thawed at room temperature in water for 1 h prior to dissection. Stringent measures were implemented to ensure the preservation of squid skin with chromatophores, minimizing the risk of any damage.
Freshly caught Indian squid samples were meticulously prepared by placing them on a dissection board. Utilizing fine-point stainless steel scissors, the posterior side of the squid was precisely cut from the base of the ventral mantle to the fin region. Subsequently, stainless steel forceps were employed to detach and remove all visceral contents, followed by cutting connective tissues to separate the visceral content from the main body. The extracted visceral content was placed in specimen bags for disposal. The remaining mantle was secured on the dissection board with stainless steel T-pins, positioning the fin side upward and submerging it in sterile chilled 0.6 M NaCl solution. The next step involved using stainless steel forceps to remove the opaque-coloured epidermal layer, which was transferred into a specimen bag, while retaining the lower chromatophore layer. After the complete removal of the epidermal layer, the chromatophore-containing skin layer was meticulously dissected into small sections measuring 1 × 1 cm. These dissected sections were then carefully transferred into 1.5 mL microcentrifuge tubes using stainless steel scissors and forceps until the tubes were halfway filled. Collected an ample number of microcentrifuge tubes, each holding a sample, and recorded the mass of every sample. Subsequently, sterile chilled 0.6 M NaCl solution was added to each microcentrifuge tube using micropipettes to final ratio of 1:1 sample to solution ratio (w/w), and the samples labelled and left overnight at +4°C for further processing. Before initiating each experiment, samples were retrieved from storage, subjected to centrifugation (Remi Laboratory Instruments, Mumbai, India) at a relative centrifugal force (RCF) of 15,124.8×g for 5 min at +4°C. The top layer of the solution was decanted and discarded into a plastic waste bucket. Subsequently, homogenization buffers were added to each microcentrifuge tube with known weights, to final ratio of 1:1 sample to solution ratio.
Entire recovery process grouped into two experimental setups. First experimental setup involved isolation of collagen from chromatophore pigment tissues, and second experimental setup included extraction of ommochrome from chromatophore pigment granules.
Measured and transferred 5 mg of pepsin and/or 30 mg of trypsin to a 50 mL screw-capped polypropylene centrifuge tube. Subsequently, added 10 mL of homogenization buffer to different test tubes containing enzymes under different buffer standardisation conditions.
Buffer conditions were meticulously defined within the homogenization buffer, and no squid samples were introduced during this phase. In this study, we explored the influence of NaCl concentration (ranging from 0% to 9%), buffer temperature (ranging from 5°C to 55°C), and buffer pH (ranging from 1 to 9), on enzyme activity. Both pepsin (European community number [EC] 3.4. 23.1) and trypsin (EC 3.4.21.4) activities were evaluated by altering conditions for each factor, while keeping all other parameters constant in the homogenization buffer (temperature = 35°C and pH = 2 for pepsin, temperature = 35°C and pH = 8 for trypsin) after vortexing at 805×g. The specific activity of pepsin or trypsin is quantified in terms of U/mg of total protein.
Selection of significant parameters and their levels by one-factor-at-a-time during recovery of collagen
In the first phase of the experiment, OFAT analysis was employed to systematically explore the influence of homogenization conditions on the recovery of collagen from chromatophore pigment tissues. The variables under investigation encompassed exposure to ultrasound (A, in minutes), temperature (B, in degrees Celsius), and pH (C) (Table 1). The impact of each variable on the extraction efficiency of collagen from chromatophore pigment tissues was determined while maintaining consistency in other parameters and conditions (temperature = 35°C, ultrasonication exposure = 30 min at 40 kHz, and pH = 2). Collagen recovery was assessed through characteristic ultraviolet-visible (UV-VIS) absorption spectral analysis using a double beam spectrophotometer (Model LMSP-UV1000B, Labman Scientific Instruments, India) at a wavelength of 230 nm. All experiments were conducted in triplicate. The centre point values, representing the parameter levels at which maximum absorption was achieved, were selected to optimize the experimental process.
| Parameter | Notations | Test range |
|---|---|---|
| Exposure to ultrasound (min) | A | 5–30 |
| Temperature of the buffer (°C) | B | 5–55 |
| pH | C | 1–9 |
The isolation of collagen from chromatophore pigment tissues was carried out in three experimental subsets.
In the first segment of the first experimental condition, a predetermined quantity of tissues underwent ultrasonication at 40 kHz for durations of 5, 10, 15, 20, 25, or 30 min by maintaining other factors at fixed value (temperature of the buffer = 35°C, pH = 2).
In the subsequent portion of the first experimental condition, a known quantity of tissues underwent ultrasonication at 40 kHz in a homogenisation buffer of pH 2 for 30 min at various temperatures (5°C, 15°C, 25°C, 35°C, 45°C, or 55°C).
In the final segment of the first experimental condition, a specified quantity of tissues underwent ultrasonication using buffers with varying pH levels (ranging from 1 to 9) at a frequency of 40 kHz for 30 min at 35°C.
The resultant homogenates were subjected to centrifugation at a RCF of 15,124.8×g for 5 min at +4°C. Subsequently, all supernatants were decanted for absorption spectral analysis at 230 nm using a UV-VIS spectrophotometer, while the infranatant containing chromatophore pigment granules was reserved for subsequent experiments.
Selection of significant parameters and their levels by one-factor-at-a-time during recovery of ommochromes
The investigation involved a systematic investigating the influence of extraction conditions on the recovery of ommochromes from chromatophore pigment granules. The extraction conditions examined included ultrasound exposure (A, in minutes), temperature (B, in degrees Celsius), and pH (C) (Table 2).
| Parameter | Notations | Test range |
|---|---|---|
| Exposure to ultrasound (min) | A | 5–30 |
| Temperature of the buffer (°C) | B | 35–85 |
| pH | C | 1–9 |
Chromatophore-containing infranatant, obtained from prior experiments, was stored in 1.5 mL microcentrifuge tubes with an equal volume of chilled 0.6 M NaCl solution following proper labelling. Subsequently, samples were withdrawn, centrifuged at a RCF of 15,124.8×g for 5 min at +4°C, and the top layer was discarded, while the bottom layer was retained for further experimentation. The extraction of ommochrome from chromatophore pigment granules was carried out in three experimental subsets.
In the first segment of the second experimental condition, the influence of ultrasound exposure durations (5, 10, 15, 20, 25, or 30 min at 40 kHz) on ommochrome extraction was examined at a pH of 2 and a temperature of 35°C.
Second segment of the second experimental condition explored the effect of temperature (35°C, 45°C, 55°C, 65°C, 75°C, or 85°C) on ommochrome extraction. This was conducted at 40 kHz for 30 min and a pH of 2.
Third subset of the second experimental configuration, we examined the influence of methanol acidity by varying the concentrations (0.010, 0.075, 0.550, 4.150, or 30.75 mL of concentrated HCl per 1,000 mL of cold methanol), resulting in pH levels of 1, 3, 5, 7, or 9, respectively, to explore its impact on ommochrome extraction. This process involved ultrasonication at 40 kHz for 30 min and a temperature of 35°C.
The resulting homogenates underwent centrifugation at an RCF of 15,124.8×g for 5 min at +4°C. All supernatants were decanted and subjected to absorption spectral analysis using a UV-VIS spectrophotometer at 280 nm.
The absorbance profiles of isolates at various recovery stages were determined using a double beam spectrophotometer within the wavelength range of 190–540 nm. Samples, obtained under specified homogenization and extraction conditions, were placed in quartz cells with a 1 cm path length. The spectrum was obtained by scanning wavelengths from 190 to 540 nm at laboratory temperature. Respective homogenization buffers or solvent served as blanks to measure absorbance during the homogenization or extraction process.
Samples collected during homogenization or extraction experiments were assayed in triplicates to determine the activity of pepsin and trypsin in the buffer. Protein, fat, moisture, and ash content of the samples were estimated as per Association of Official Analytical Chemists International (AOAC) (1999) method. The specific activity of both pepsin and trypsin is expressed in U/mg of total protein.
The activity of pepsin in the homogenization buffer was measured as trichloroacetic acid-soluble product using haemoglobin as the substrate through stop rate determination (Anson, 1938), employing a spectrophotometer with a light path of 1 cm at A280. One unit of pepsin produces a ΔA280 of 0.001/min with haemoglobin as the substrate at pH 2.0 at 35°C in a 3.2 mL sample. The enzyme unit/mL is calculated as ([A280 test – A280 blank] × [df])/([10] × [1.0] × [0.001]), where df is the dilution factor, 10 is the assay incubation time in min, 1.0 is the volume of enzyme solution (mL) added, and 0.001 is the ΔA280 per unit of pepsin.
The activity of trypsin in the homogenization buffer was measured using Nα-Benzoyl-L-arginine ethyl ester (BAEE) as the substrate through continuous rate determination (Bergmeyer, 1974), employing a spectrophotometer at A253 with a light path of 1 cm. One BAEE unit of trypsin activity produces a ΔA253 of 0.001/min with BAEE as the substrate at pH 8.0 at 35°C in a sample volume of 3.2 mL. The BAEE enzyme unit/mL is calculated as ([ΔA253/min test – ΔA253 blank] × [df])/([0.001] × [0.075] × [0.001]), where df is the dilution factor, 0.001 is the change in A253/min based on unit definition, and 0.075 is the volume (mL) of the test sample used in the assay.
In this study, the central composite design (CCD) is utilized as a fractional factorial design within the response surface model.
Utilizing CCD, RSM proves valuable as a mathematical and statistical approach for experiment design. It enables the analysis of interactions among independent variables and facilitates the creation of an optimal synergy of responses prior to assessing response patterns (Khuri & Mukhopadhyay, 2010).
Two independent factors, namely exposure to ultrasound in minutes and the temperature of the buffer in degrees Celsius, were investigated across five levels: −α (minimum), −1 (low), 0 (central), +1 (high), and +α (maximum). These levels were chosen based on the results of the OFAT study. Thirteen experiments were conducted to analyse data and assess the interaction between two independent variables across five levels of process efficiency. The software Design Expert (StatEase, Minneapolis, MN, USA) was employed for this analysis.
RSM was employed to optimize the sequential recovery process, initial stage focusing on extracting collagen from chromatophore pigment tissue while leaving behind chromatophore pigment tissue granules. Subsequently, the extraction process targeted ommochrome from chromatophore pigment tissue granules. The key variables identified as significant for optimization were exposure to ultrasound (A) and temperature of the buffer (B). These variables were incorporated into a CCD while maintaining a constant pH of 2 during the first stage.
Moving to the second stage, the significant variables for optimization were still exposure to ultrasound (A) and temperature of the solvent (B). These variables were integrated into a CCD, and a fixed pH of 1 was maintained. Center points for the CCD were selected based on the outcomes of OFAT experiments.
To ensure robust experimental design, CCD experiments were generated and executed in a random order. The details of the experimental parameters and their corresponding levels for the first experiment is presented in Tables 3 and 4.
| Variables | Notations | Levels | ||||
|---|---|---|---|---|---|---|
| –α | –1 | 0 | +1 | +α | ||
| Exposure to ultrasound (min) | A | 25.86 | 30 | 40 | 50 | 54.14 |
| Temp of the solvent (°C) | B | 45.86 | 50 | 60 | 70 | 74.14 |
The specifics of the experimental parameters and their respective levels for the second experiment for ommochrome extraction are outlined in Tables 5 and 6.
| Variables | Notations | Levels | ||||
|---|---|---|---|---|---|---|
| –α | –1 | 0 | +1 | +α | ||
| Exposure to ultrasound (min) | A | 25.86 | 30 | 40 | 50 | 54.14 |
| Temp of the solvent (°C) | B | 45.86 | 50 | 60 | 70 | 74.14 |
The experimental data underwent multiple regression analyses, expressing the mathematical relationship between response values and five independent variables through a second-order polynomial model as defined by Equation (1) as follows.
Here, Y represents the process response predicted, β0 is the constant coefficient, βi represents the linear coefficient, βii signifies the quadratic effect, βij denotes the interaction effect, k is the number of studied independent factors, and xi, xj, are coded independent variables.
Once statistical modelling and regression analysis was performed, Analysis of Variance (ANOVA) was employed with a significance level (p-value ≤ 0.05) F-value and R2 to determine the significance of each factor in the fitted equations and to evaluate the adequacy of the fit in each instance (Asadzadeh et al., 2018).
Design Expert was utilized to generate two-dimensional (2D) contour plots and three-dimensional (3D) response surface plots, serving as graphical representations for the regression equation to optimize reaction conditions. This approach proved valuable in elucidating the conditions of the reaction system by illustrating the simultaneous interaction effects of two-factor levels on responses. The plots featured removal percentage (response) on the Z-axis and two independent variables on the X and Y axes, while keeping other variables at the coded zero point (Sarrai et al., 2016).
The validity of the second-order polynomial model derived from RSM was confirmed through a set of experiments, where random parameter values within the optimized levels were chosen. Additionally, experiments were carried out under the conditions optimized by the software. The experimental results were subsequently compared to the predictions made by the second-order model obtained from CCD to assess the model’s goodness of fit. The system’s behavior is elucidated through the quadratic equation presented in the Equation (1).
ML approach was adopted to enhance the reliability of the model. Experimental data obtained from the CCD (collagen and chromophore) were subjected to training, testing and prediction by ML approach (Lana, 2023) such as ANN, Random Forest, kNN using Orange Data Mining Tool (version 3.36.1, University of Ljubljana, Ljubljana, Slovenia). Workflow of the procedure is schematically depicted in the Fig. 1.
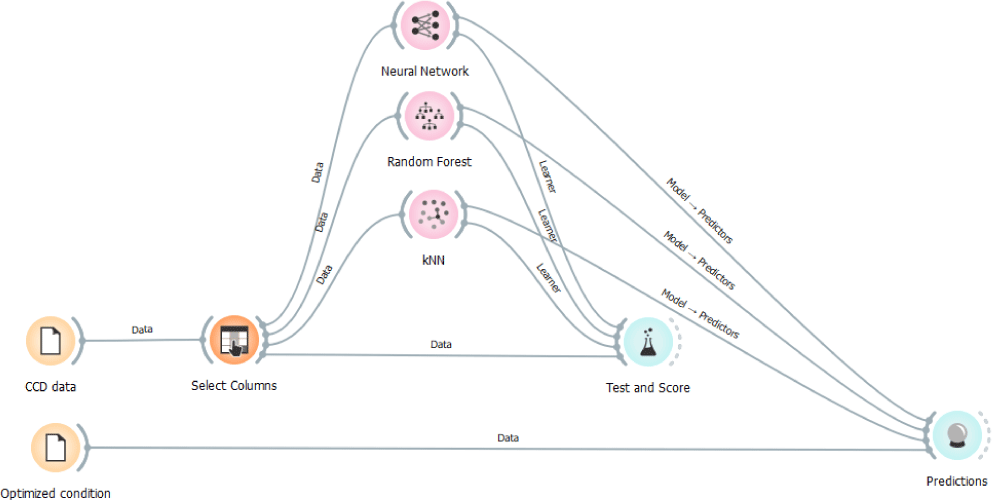
“CCD data” widget contains training and testing datasets. Exposure to ultrasound and temperature of the buffer were chosen as features and collagen as target in the “select columns” widget and connected to neural network widget for configuration. Trained model was used for prediction. Optimized conditions widget was connected to prediction widget to obtained predicted value. Chromophore CCD data was trained, tested and predicted using the same workflow.
Results and Discussion
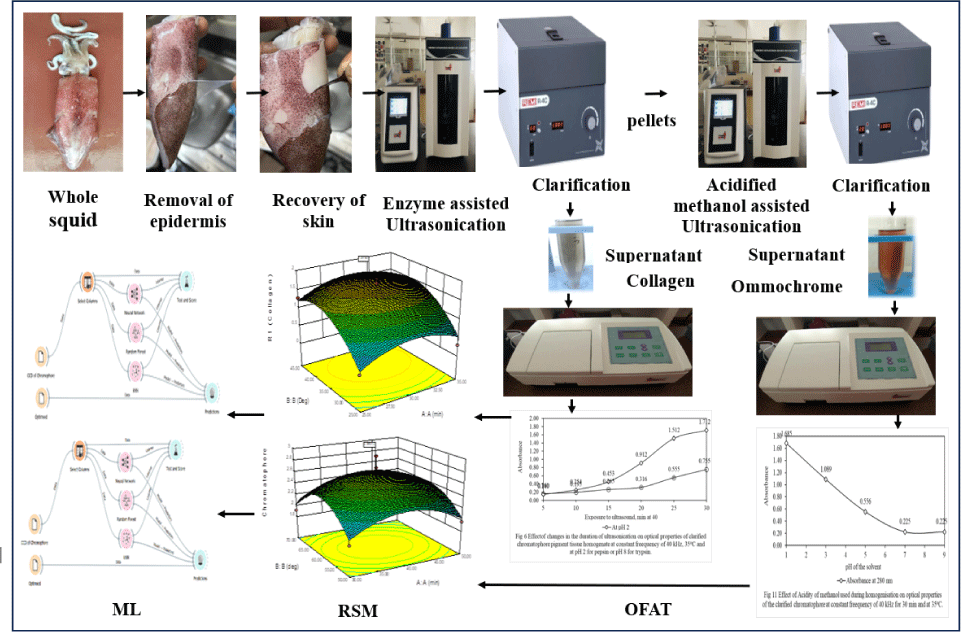
Our study effectively optimized the sequential extraction of collagen and ommochrome from Indian squid epidermal tissues, a valuable byproduct of the thriving Indian seafood industry (PIB, 2023a, 2023b). Seafood processing generates significant waste biomass (Olsen et al., 2014) rich in bioactives like collagen and chromatophores (Nam et al., 2008), which offer applications in food, pharma, and textiles due to properties like UV absorption (Lewis et al., 2022). Our approach utilizes ultrasound-assisted extraction, a prominent, sustainable method. This aligns with research showing ultrasound, sometimes combined with enzymatic hydrolysis, efficiently extracts high-value compounds. For example, Lee et al. (2022) used combined techniques for antioxidant collagen hydrolysates, while Chan et al. (2019) optimized ultrasonication for antibacterial ommochromes from squid skin. Such targeted extraction transforms marine waste into valuable resources (Chan et al., 2019; Chen et al., 2023; Crawford, 2020; Esparza-Espinoza et al., 2022). Unlike single-product, multi-stage recoveries (Lewis et al., 2022), our novel two-stage process for both collagen and ommochromes, optimized using OFAT, RSM, and advanced ML models (ANN, Random Forest, kNN), significantly contributes to sustainable marine byproduct valorization (Chauhan et al., 2021; Noi & Kappas, 2017; Reungoat et al., 2021).
In our study, we meticulously removed the chromatophore-containing skin layer from the epidermal layer, mantle, tentacles, wings, and visceral layer (Fig. 2). The Indian squid specimen exhibited a composition of 74.6 ± 0.30% moisture, 22.2 ± 0.12% protein, 1.1 ± 0.02% fat, and 1.2 ± 0.03% ash. The results are in line with the previously published research papers on Loliop lei (Lapa-Guimarães et al., 2005). Among the total protein content (22.2 ± 0.12%), 63.5 ± 0.70% comprised myofibrillar protein, 13.9 ± 0.19% sarcoplasmic protein, 13.2 ± 0.32% alkaline-soluble protein, and 12.0 ± 0.07% total collagen. Given that myofibrillar protein is water-soluble and may leach into the medium during subsequent processing, careful removal is crucial. Notably, while the mantle contains 12.0% of total collagen, the tentacles of the Indian squid also harbor 12.8%, and the epidermal connective tissue contains 17.8% collagen. Raman & Mathew (2014) reported that the squid connective tissues of epithelial cell has highest amount (17%) of collagen than the collagen content in the mantle (12%). The squid features an outer transparent epidermal layer composed of columnar epithelial cells interspersed with mucous cells, and a lower dermal layer comprising connective tissues housing chromatophores, iridophores, and reflecting cells. The epidermal layer can be effectively separated from the lower dermal layer through precise dissection using scissors. The connective tissues of the dermal layer consist mainly of fibroblasts and a network of collagen fibers (Anadón, 2019) that incorporate localized chromatophores in elastic sacs. Given the necessity to isolate ommochromes from chromatophores pigment tissues, a careful strategy was devised to minimize the initial load of collagen in the raw material, thereby facilitating the isolation of the pigments from the tissues.
The activity of pepsin and trypsin in homogenization buffer was evaluated under varying NaCl concentrations, temperatures, and pH conditions in the absence of any substrate. No significant change in pepsin activity was observed at 0% and 1% NaCl when other parameters were constant (35°C, pH 2); however, pepsin activity declined significantly at NaCl concentrations ≥ 2%, possibly due to ionic strength interfering with enzyme-substrate interactions (Mazumder et al., 2018). For trypsin, increasing NaCl concentration (from 0% to 4%) under constant temperature (35°C) and pH (8) led to a consistent drop in activity by more than 50%, suggesting a salting-out effect and enzyme precipitation due to ionic competition for water and enhanced hydrophobic interactions (Cheison et al., 2011).
Pepsin activity increased with rising temperature and peaked at 35°C, while further increases led to thermal denaturation. This aligns with Mazumder et al. (2018), who reported that human pepsin exhibits maximal activity at 37°C, with a sharp decline beyond 40°C. Similarly, trypsin showed optimal activity between 35°C and 55°C in buffer (0% NaCl, pH 8), as reported by Cheison et al. (2011), who observed trypsin retaining over 80% of its activity at 45°C and nearly complete inactivation at temperatures below 25°C.
Pepsin exhibited activity in the pH range of 1–4, with peak activity at pH 2 (at 35°C, 0% NaCl), and its activity sharply decreased beyond pH 5. Mazumder et al. (2018) confirmed that pepsin is irreversibly denatured above pH 6.5. Trypsin activity was optimal at pH 8 and functional in the range of pH 7–9, with marked loss of activity below pH 6 due to irreversible conformational changes and denaturation (Klomklao et al., 2012). Klomklao et al. (2012) demonstrated that trypsin from marine sources lost over 70% activity at pH 6 and was inactive at pH 5.
Based on these findings, a homogenization buffer incorporating both pepsin and trypsin was formulated at 35°C, pH 2, and without added NaCl to maintain optimal activity and structural integrity of both enzymes during the extraction process.
We investigated how individual homogenization parameters influenced the yield and optical properties of clarified collagen homogenates at 230 nm, keeping all other variables constant. To isolate collagen from the dermal connective tissues that is rich in fibroblasts and collagen networks, we applied a combination of enzymatic treatment and ultrasound exposure. After homogenization, we centrifuged the samples at 15,124.8×g for 5 minutes at +4°C, which effectively separated the collagen-rich supernatant from pigment-laden chromatophore pellets.
The presence of collagen in the supernatant was confirmed by specific UV absorbance at 230 nm, indicative of glycine, proline, and hydroxyproline—amino acids characteristic of the collagen triple helix. This method yielded 69.69% collagen from epidermal tissues. For comparison, Tan & Chang (2018) reported a 64.19% recovery using pH 2.4 HCl and 23.6 KU/g pepsin from catfish skin, noting smaller α-chains and higher gel strength in ASC and HSC than PHSC, underscoring the industrial potential of homogenization-aided acid extraction.
Next, we assessed how ultrasound exposure time, homogenization temperature, and buffer pH affected collagen release from chromatophores. In the first condition, the impact of individual homogenization factors on the optical properties of the clarified homogenate at 230 nm was evaluated, with other variables held constant (Fig. 3). Traditional enzyme-based methods, with or without acids, yield low collagen from tough dermal tissues due to matrix resistance. Hence, the current investigation focused on specific factors such as ultrasound duration, temperature, and buffer pH, to identify significant contributors to collagen isolation (DiBona et al., 2016; Lee et al., 2022; Oancea et al., 2012). Protein UV absorbance is shaped by structural features such as peptide bonds and side chains. Collagen’s triple helix has a defined absorption peak at 230 nm due to its unique amino acid content (Kittiphattanabawon et al., 2015).
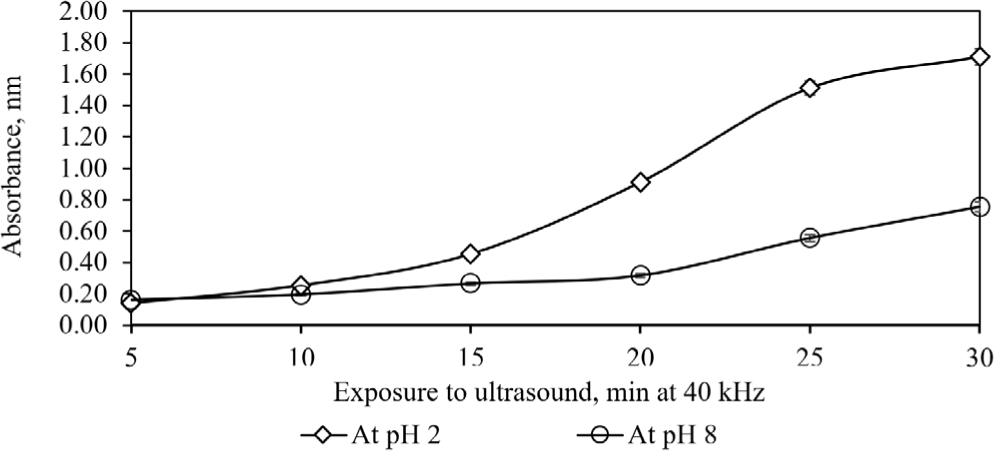
An increase in ultrasound exposure enhanced absorption at 230 nm. Ultrasonication, a non-thermal technique, proved effective in improving collagen yield, reducing extraction time, and outperforming either enzyme or ultrasound alone. Enzyme-assisted ultrasonication showed superiority, especially at buffer pH 2, doubling yield compared to pH 8. Pepsin and trypsin were effective in collagen extraction from squid skin. Varying pH while keeping exposure time constant (e.g., 30 min) is fundamental to establish cause-effect relationships in experimental design (Montgomery, 2019).
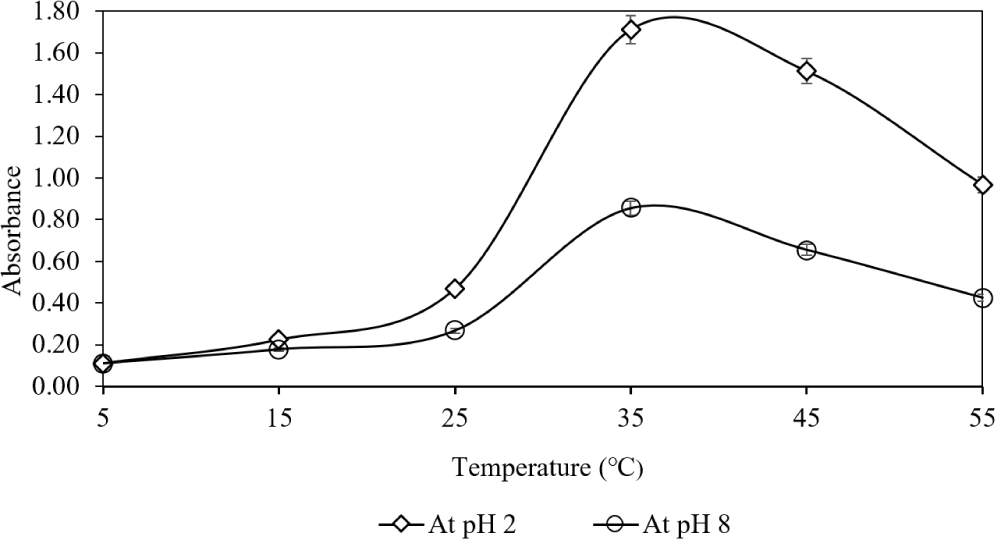
The second part of this condition evaluated the role of temperature during enzyme-assisted ultrasonication on clarified tissue absorbance at 230 nm (Fig. 4). Increasing temperature to 35°C enhanced absorbance, but further increases led to decline. Literature suggests pepsin homogenization preserves the collagen helix, with denaturation starting near 35°C (Jafari et al., 2020).
In the third phase, we studied how buffer pH during enzyme-assisted ultrasonication influenced clarified homogenate properties (Fig. 5). Holding other parameters constant, two absorption peaks were observed. The major peak at pH 2 aligned with pepsin activity, while a smaller peak at pH 8 indicated trypsin activity, consistent with Feng et al. (2013).
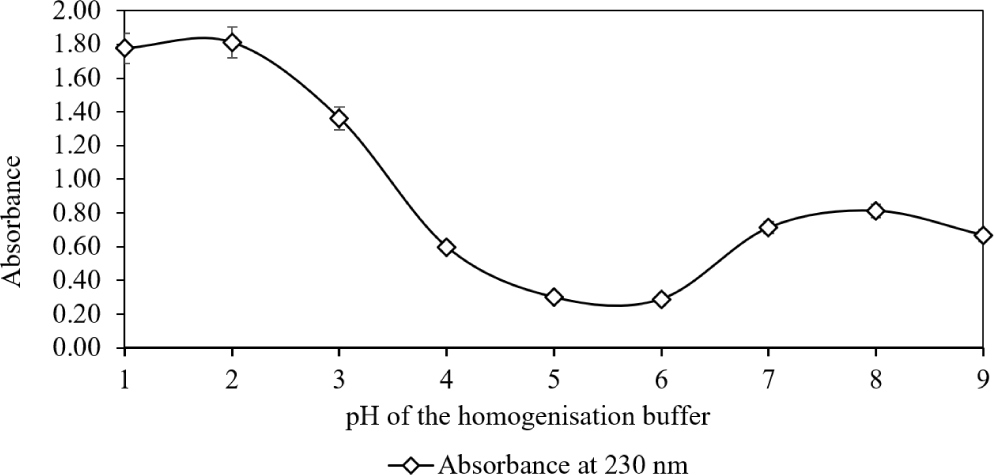
Overall, enzyme-assisted ultrasonication significantly enhanced collagen extraction. Ultrasound duration, homogenization temperature, and buffer pH each played key roles. Increasing ultrasound exposure facilitated matrix disruption, and temperature control was critical, with 35°C as the upper threshold before collagen denaturation. Buffer pH influenced enzyme performance, pepsin was most active at pH 2, outperforming trypsin near pH 8. These findings establish that a combined method involving homogenization, enzymatic digestion, and ultrasound optimally extracts collagen. Operating at pH 2, 35°C, and a suitable ultrasound duration yields a scalable, efficient strategy for recovering collagen from chromatophore-rich tissues.
Our research offers a unique, optimized strategy for extracting collagen from complex dermal tissues, particularly emphasizing how enzyme-assisted ultrasonication, precise temperature control (up to 35°C), and specific pH levels (optimally pH 2 for pepsin) dramatically enhance yield and overcome matrix resistance where traditional methods fall short. Unlike previous studies, we pinpoint the critical interplay of these factors to establish a highly efficient and scalable recovery process for collagen, specifically from chromatophore-rich tissues. This optimized methodology directly supports the industrial demand for high-quality collagen, enabling its sustainable extraction for diverse applications from challenging biological sources.
The pellets from the initial experiment were then used in a second study to investigate the impact of ultracentrifugation, extraction temperature, and methanol acidity on ommochrome extraction from chromatophore sacs (Chan et al., 2019). The first part of this second study focused on how ultrasound exposure during homogenization affected the optical properties of clarified chromatophores, keeping other variables constant (Fig. 6). Increased exposure led to higher ultraviolet absorption in the extract. We found a chromatophore yield of 12.1% from epidermal tissue, with a 1.24% yield of soluble pigment from these chromatophores. These results align with DiBona et al. (2016), who reported 2–3 mL of soluble pigment per squid Doryteuthis pealeii sample from HCl-MeOH extraction, representing 58% of the yield from pigment granules.
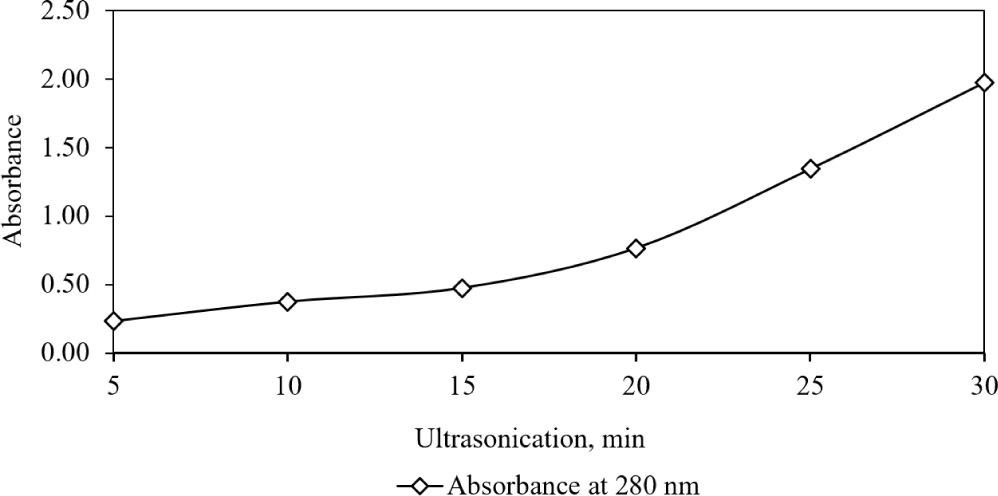
The second segment of the second experimental condition studied the impact of temperature of the homogenisation on the optical properties was investigated while maintaining other conditions constant. UV-VIS absorption spectral analysis of the supernatant, extracted using HCl-MeOH at temperatures ranging from 35°C to 85°C, clearly demonstrated a significant increase in absorption magnitude up to 65°C (Fig. 7). However, a notable decline in the absorption peak was recorded at 75°C. This aligns with findings by Chan-Higuera et al. (2019), who reported increased pigment yield (635 mg/g from Dosidicus gigas skin) with rising ultrasonication time (5 to 15 min) and temperature (25°C to 45°C). However, they also observed a decrease in the antimicrobial and antimutagenic activities of the extracted pigment as temperature increased from 25°C to 45°C.
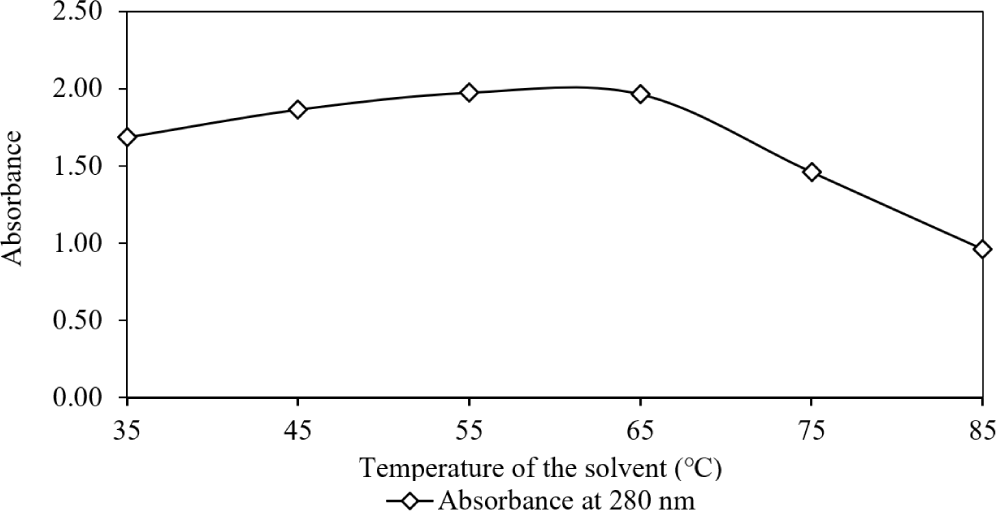
Moving to the third segment of the second experimental condition, the impact of changes in acidity of methanol during homogenization on the photo absorptive characteristics of clarified chromatophores was determined while holding other factors constant (Fig. 8).
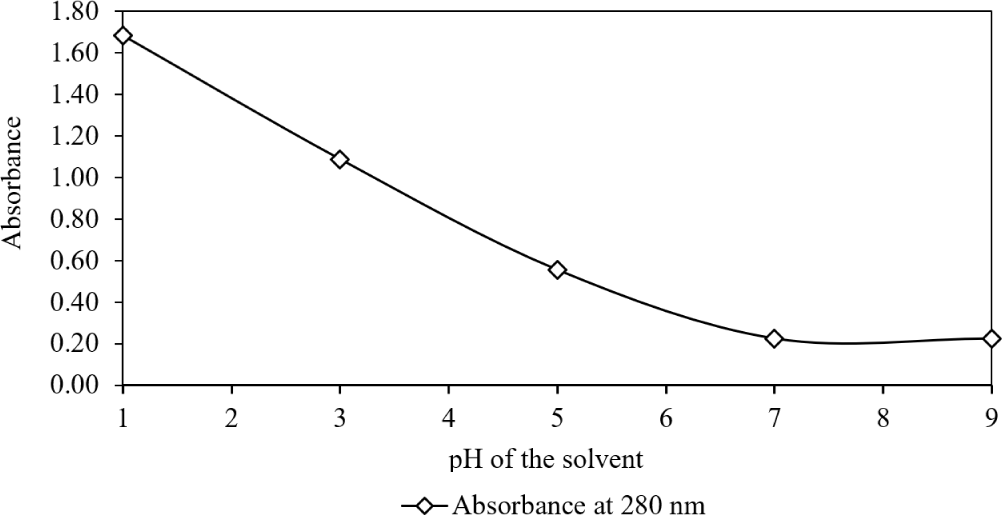
Elevated methanol acidity significantly enhanced ommochrome extraction, evident through an increase in absorption. This aligns with existing research supporting the efficacy of HCl in methanol for pigment extraction (DiBona et al., 2016; Simonian, 2002). Analysis of the extract at 280 nm was conducted to monitor proteinaceous components (Simonian, 2002) and, indirectly, the presence of chromatophores, as ommochromes and associated proteins within chromatophores exhibit absorption in the UV region (Williams et al., 2016).
Our research uniquely advances ommochrome extraction by systematically optimizing parameters like ultrasound exposure, temperature, and methanol acidity, providing specific thresholds for maximizing yield while considering pigment stability. Unlike broader studies that focus solely on extraction efficiency, we highlight the critical balance needed to prevent thermal degradation that could compromise the extracted ommochromes’ inherent properties. These precise optimization insights are directly relevant to industries seeking efficient, high-quality biopigment recovery, opening avenues for sustainable natural colorants and high-value biochemicals.
The UV-VIS absorption spectra of collagen and ommochromes recovered from squid skin were analysed. The supernatant displayed its maximum absorption at 230 nm, as illustrated in Fig. 9. Notably, in our investigation, the supernatant exhibited no absorption peaks at 280 nm. Typically, proteins containing amino acids like tyrosine and phenylalanine manifest absorption peaks at 280 nm (Biter et al., 2019). Our findings propose that the lack of discernible absorption peaks within the spectrum of 250 to 280 nm indicates the absence of proteins with aromatic amino acids, such as phenylalanine and tyrosine, in the supernatant. These amino acids are crucial components of sensitive chromatophores (Reinmuth-Selzle et al., 2022).
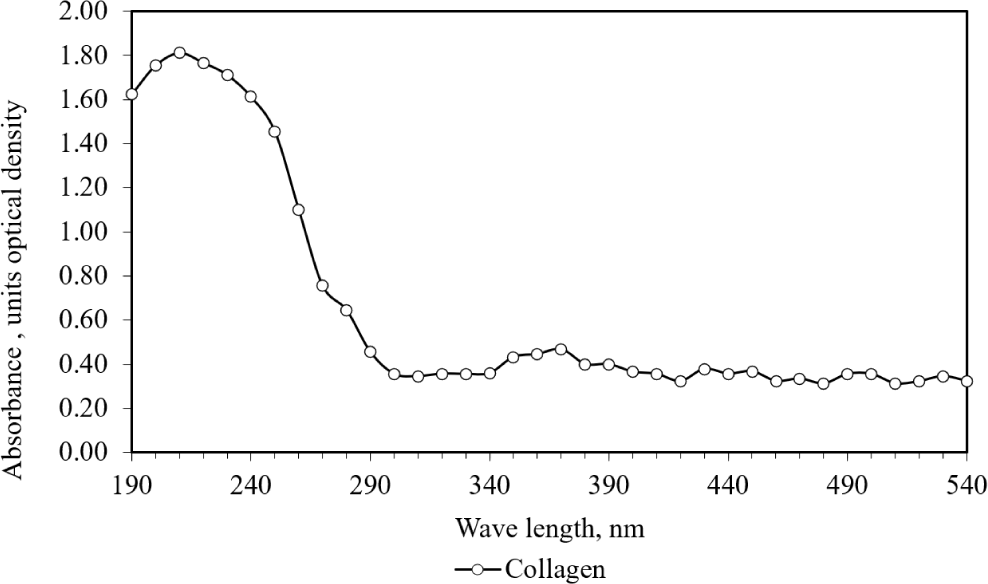
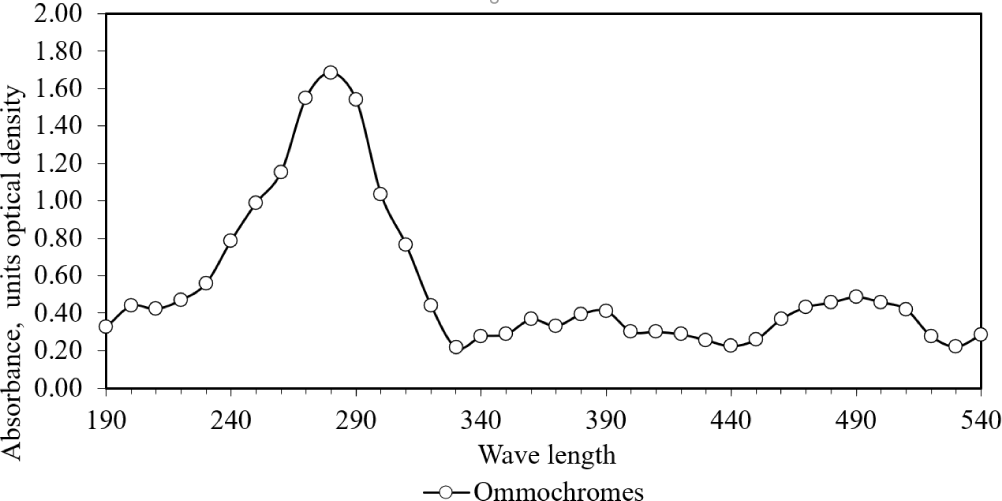
The absorption of UV-VIS rays by the supernatant, obtained through HCl-MeOH extraction and subsequent clarification via centrifugation, revealed distinctive absorption peaks at 280 nm, along with smaller peaks at 370 and 470 nm (Fig. 10). Additionally, the purity of the chromatophore pigment granule constitutes a significant factor that could potentially influence the efficiency of HCl-MeOH extraction (Szymborska et al., 2022).
All experiment of collagen recovery were designed using CCD to assess interactive effect of significant factors on response using RSM. Significant factors influencing collagen recovery, identified through OFAT experiments, include exposure to ultrasound (A, in min) and temperature of the buffer (B, in degree Celsius). Variables, A and B were tested at five different levels –α (minimum), –1 (low), 0 (central), +1 (high) and +α (maximum) obtained from preliminary Experiments conducted, keeping all other factors constant. Design expert recommended 13 experimental runs, comprising 4 factorial points, 4 axial points, and 5 center points, to enhance collagen recovery. Using the proposed model, a quadratic equation was formulated to forecast the dependent variables A and B by considering the independent variables and their interactions. Experiments were performed under different conditions as per the experimental run suggested by CCD. A second-order polynomial equation, derived from experimental results, illustrates the interactions among the proposed independent variables A and B in relation to collagen recovery efficiency. It is expressed as follows:
The model’s adequacy was evaluated through ANOVA applied to experimental data. The importance of each coefficient in the model was assessed through the F-test and probability of error test (p-value) for ANOVA, conducted at a 95% confidence level. Model terms were considered significant when the p-value was less than 0.005 and deemed not significant when p-values exceeded 0.1. Table 7 presents the results of the ANOVA on this model, demonstrating its high significance with a regression p-value of 0.0004. Here, the total variation of results is partitioned by ANOVA into components related to models and components related experimental errors, which in turn helps in determining the significance of the model’s variation (Yusuff, 2018). The calculation involves evaluating the F-value, expressed as the square-to-residual error ratio of the mean model. If the calculated F-value surpasses the tabulated F-value, the model proves to be a robust predictor of experimental data (Ani et al., 2015). In the current investigation, the obtained F-value was 21.63, indicating the fitness of the response surface model. Additionally, the significance of each model term was assessed using the probability of error value (Prob > F). Values in Table 7 below 0.0004 suggest the significance of the terms (Garba et al., 2016). Notably, B, A2, and B2 were deemed significant model terms, while A and AB were considered insignificant for collagen. The F-value indicates that the two studied process variables significantly affected the yield of collagen. The highest F-value, 54.02, implying that temperature had the most substantial influence on collagen yield compared to exposure to ultrasound. However, only the quadratic effects of exposure time (A) and temperature (B) on collagen release were found to be significant.
The lack-of-fit test coefficient was introduced to assess whether the chosen model adequately described the observed data or if a more complex model should be considered. Table 7 reveals a lack-of-fit F-value of 2.92 and p-value of 0.1639, indicating that the lack of fit is not statistically meaningful relative to pure error. A non-significant model inadequacy is advantageous because the model aims for perfect fitness. The second-order polynomial model not only proved to be statistically significant but also demonstrated an insignificant lack of fit. Here, the p-values are greater than 0.05 for the model interaction (p-value: 0.1639 > 0.05). This suggests that the polynomial second-order model corresponds closely to the experimental collagen extraction data. The determination coefficient (R2) for the model was calculated to be 0.93, indicating a high level of explanatory power. When R2 values approach unity and standard deviation values are smaller, it signifies a more accurate predictive capability of the developed model (Chicco et al., 2021).
The second-order polynomial models describing collagen and chromophore extraction, respectively. Table 8 displays the results of fitting the quadratic response surface model, presenting the ANOVA outcomes.
Interactive effect of exposure to ultrasound (A) and temperature of the buffer (B) and optimum levels of these variables on the synergy of absorbance at 230 nm due to collagen is illustrated using 2D contour graphs (Fig. 11) and 3D surface plots were used (Fig. 12) based on the regression equation. RSM was employed to establish a relationship between exposure to ultrasound (A) and temperature of the buffer (B), and UV absorptive properties of collagen. Maximum absorbance value of 1.812 due to collagen was obtained at 30.453 min of ultrasound exposure and a temperature of 38.80°C, and the desirability is 0.936. The UV absorption at 230 nm increased with ultrasound exposure and temperature, displaying significance when the contour plot shape was elliptical. Contour plots of elliptical shape as shown in Fig. 11 indicates that the interplay among corresponding variables, A and B is significant. The concave shape of the response surface indicated significance (El-Mekkawi et al., 2021), supporting collagen extraction. In this study, the observed response levels in the experiment align with the theoretically assumed values of the model, demonstrating the precision and accuracy of the response surface models.
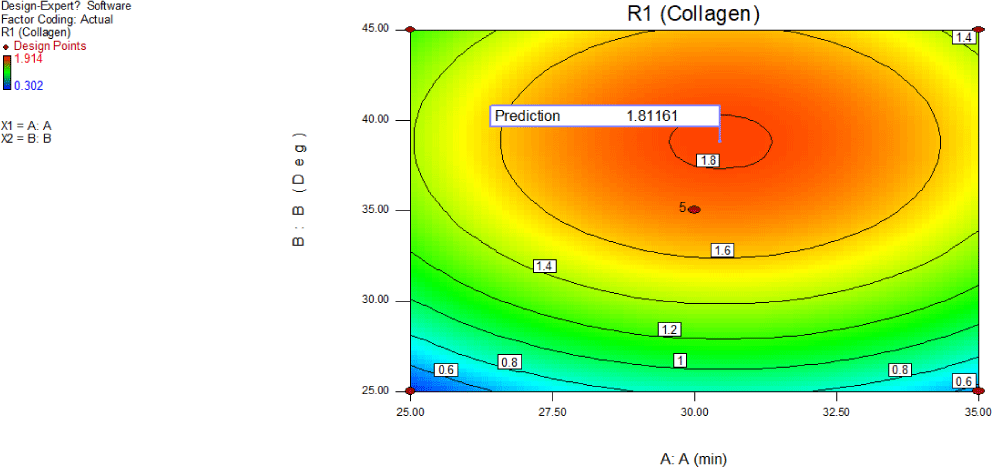
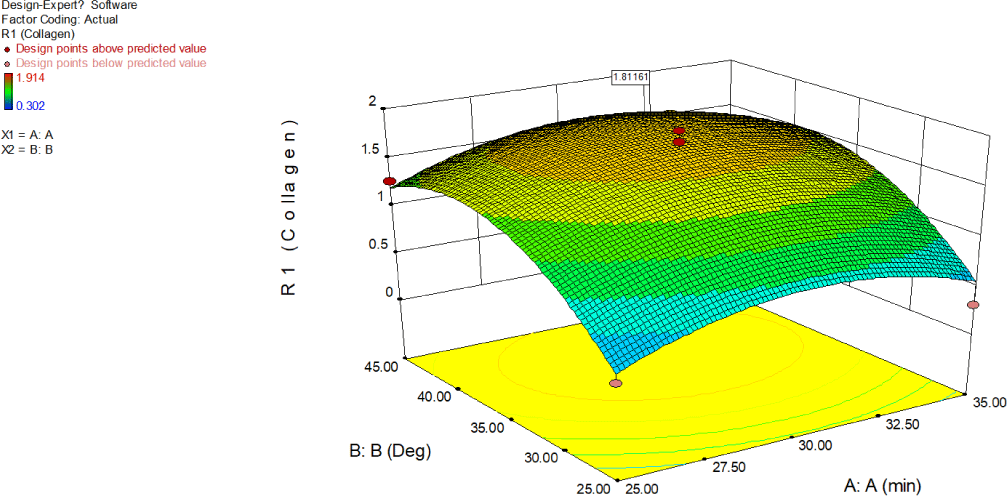
An increase in exposure to ultrasound (A) and temperature of buffer (B) positively influenced the absorbance at 230 nm due to collagen. Consequently, exposure to ultrasound (A) and temperature of buffer (B) exhibited the most significant positive impact on the absorbance at 230 nm due to collagen. Conversely, an increase exposure to ultrasound (A) and temperature of buffer (B) led to a decrease in absorbance at 230, indicating negative effects on the recovery process. The curvature in the Fig. 12 exhibited by exposure to ultrasound and temperature of buffer endorse significance on collagen extraction.
Within response surface designs, the perturbation plot illustrates how the response varies as each factor deviates starting from the selected reference point, while keeping all other variables constant at their reference values (Fig. 13). A perturbation plot was generated to better identify the most influential factors in collagen recovery. Specifically, the perturbation plot for collagen recovery was utilized in this context to evaluate behavior of the response when deviating from the midpoint, while maintaining the constant values of other factors. Derived from the perturbation plot, conclusion can be drawn that both variables exposure to ultrasound and temperature significantly influenced collagen extraction.
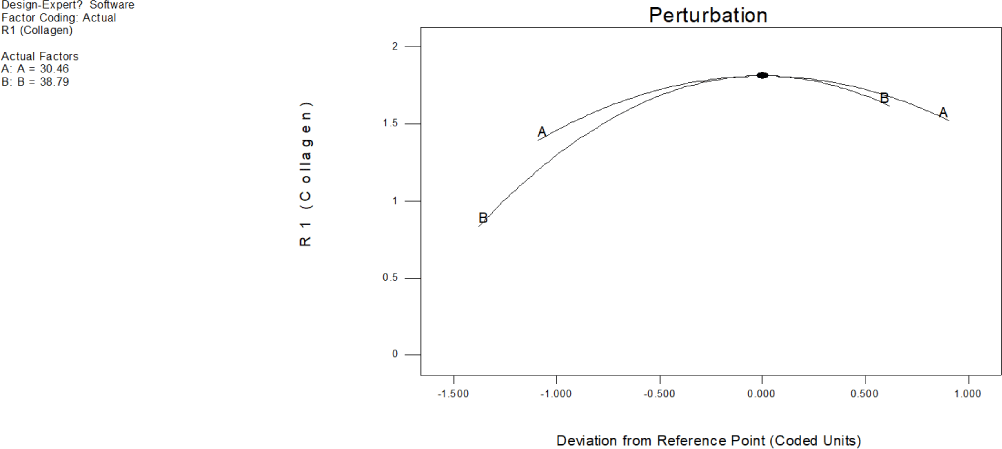
Fig. 14 illustrates yield plot of collagen recovery reveals the spots distributed uniformly along the diagonal axis. Linear graph that intersects the origin suggest a close alignment between actual experimentally observed collagen yield and the predicted values derived through optimisation methodology (Saeed et al., 2015). The scatter plot of residuals against increasing predicted responses conforms to the principle of constant variance, showcasing a uniform spread of residuals across the entire graph. The experimental findings affirm the suitability of the quadratic model, obviating the necessity for further transformations of the data series (Bavisetty et al., 2024; Sadowska & Sikorski, 1987).
Two-factor and five-level CCD guided by Design Experts was used to conduct 13 runs, incorporating factorial, axial, and center points to optimize ommochrome recovery. RSM was used to model experiments to predict A and B in terms of independent variables. Second order polynomial model developed for the second experiment is as follows:
The adequacy of the model was assessed using ANOVA applied to experimental data. The F-test and probability of error test (p-value) for ANOVA were utilized to assess the significance of each coefficient in the model. Table 9 displays the ANOVA results, indicating a regression p-value of 0.0042. In this context, a p-value below the significance level of 0.05 suggests that the experimental data on ommochrome recovery offers ample evidence to assert that the regression model fits the experimental data more effectively than the model incorporating independent factors. F-value of the 10.11 indicates the robustness of the response surface model as a predictor of experimental data.
High F-value of 10.11 and p-value of 0.0042 indicates that the regression robustness of the model (Karchiyappan & Karri, 2021). Experimental results were partitioned by ANOVA into model-related components and experimental error-related components. Notably, A2 with the highest 37.12, and B2 were with the value of 19.42 deemed significant model terms, suggests that the two studied process variables significantly affected ommochrome yield. The F-value suggests that the two studied process variables significantly affected ommochrome yield, with the highest F-value (37.12) affiliated to B2, indicating temperature’s substantial influence on ommochrome yield compared to exposure to ultrasound. However, only the quadratic effects of exposure time (A) and temperature (B) on chromophore release were found to be significant.
Lack-of-fit test coefficient was introduced CCD under RSM to assess the descriptive capabilities of model. Table 7 reveals that ANOVA of the experiment of ommochrome recovery has the p-value of 0.6067 which is greater than 0.05 for the model. In RSM, the p-values greater than 0.05 for the model interaction indicate that the polynomial second-order model aligns well with the experimental ommochrome extraction data and it is advantageous. The determination coefficient (R2) for the model was calculated to be 0.76. High R2 values approaching unity, coupled with smaller standard deviation values, indicating that the associated predictions are very close to experimental condition for the recovery of the ommochrome. Here, analysis of ANOVA confirms that the second-order quadratic model is superior in predicting the CCD of ommochrome recovery.
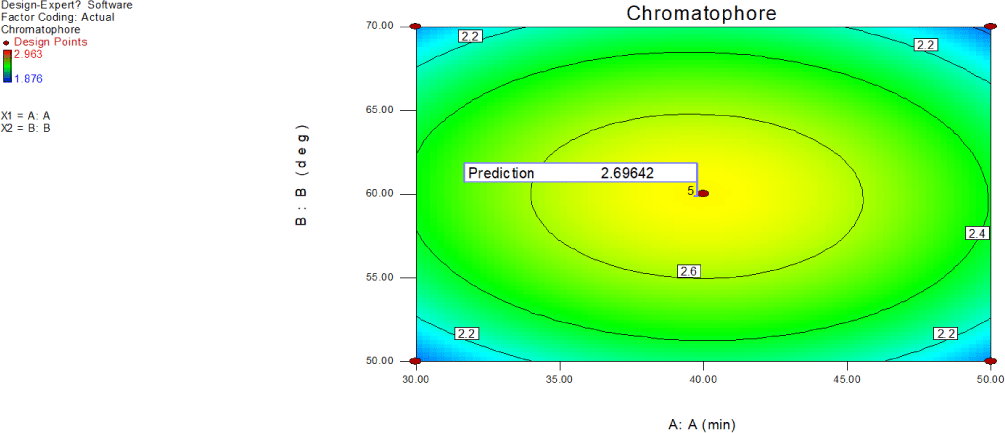
Figs. 15 and 16 graphically depict the 2D and 3D plots illustrating the absorbance at 280 nm, attributed to ommochrome, based on the concerted impacts of ultrasound exposure duration (A) and solvent temperature (B). The enhancement of working conditions was achieved through RSM to maximize system performance. The maximum absorbance value of 2.6964, attributed to ommochrome, was achieved at 39.78 min exposure to ultrasound and a temperature of 59.87°C. The UV absorption at 280 nm showed a notable rise with increased ultrasound exposure and temperature, especially in instances where the contour plot displayed an elliptical shape. The elliptical contour plots signify the significance of the inter effect among respective variables A and B. The convex shape of the response surface further indicated significance, providing support for ommochrome extraction. The quadratic model was statistically meaningful, and inadequate fitting was insignificant (p-value: 0.6067 > 0.05). It signifies that polynomial second order model fits well the experimental collagen extraction data. The model’s determination coefficient (R2) stood at 0.87.
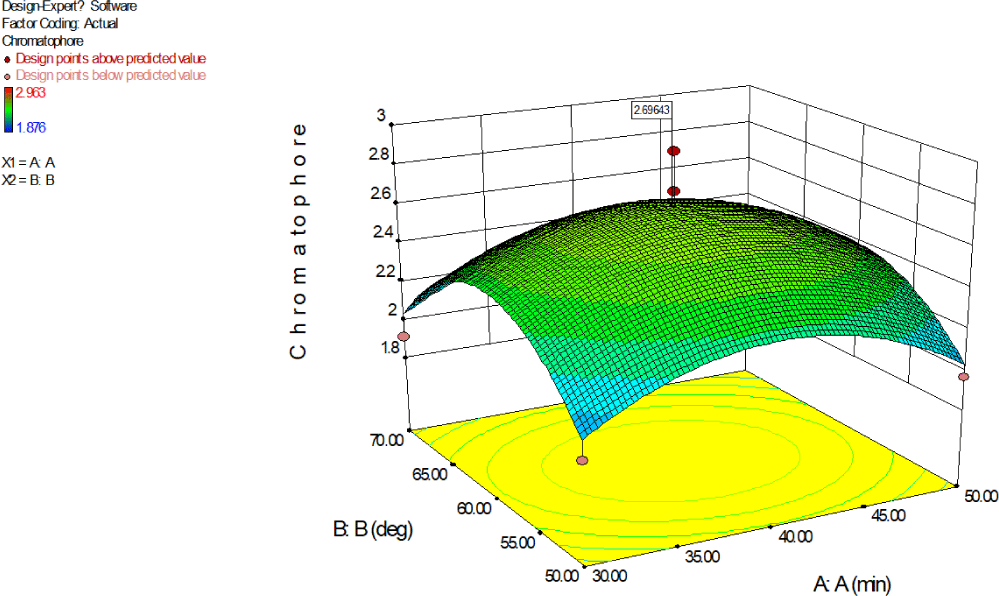
Here, shape of the curve opened downward. Increase in the exposure to ultrasound and increase in temperature increased the absorption due to ommochrome at 280 nm, however continued exposure to ultrasound and continued increase in reaction temperature decreased the absorption due to ommochrome at 280 nm. The desirability value achieved was 0.755, which remains within an acceptable range (Sai-Ut et al., 2023) for methanol extraction.
The experimental outcomes and anticipated values were 2.739 and 2.696 nm, respectively. These results showcase a robust concordance between the observed and forecasted values under the optimal operational conditions. Thus, the regression models obtained by RSM accurately predict the (Aydar et al., 2017) absorbance at 280 due to ommochrome for various combinations of ultrasound exposure and solvent temperature.
The composite desirability values, representing the overall quality of optimal conditions, were 0.755. This value, considering multiple factors and responses, indicate how well the chosen conditions perform in achieving desired outcomes for ommochrome. A higher desirability value suggests that the conditions are more favorable for maximizing extraction yield while minimizing extraction time or solvent usage. In this instance, the conditions for ommochrome achieved higher desirability values, indicating a more optimal parameter set for the extraction process (Butenandt et al., 1957; Chan et al., 2019; Chen et al., 2023; Esparza-Espinoza et al., 2022). The experimental data align with the predictions, affirming the validity of the model used and the efficiency of RSM in optimizing conditions for extracting ommochrome from chromatophore sac.
An increase in exposure to ultrasound (A) and the temperature of the methanol (B) had a favorable effect on absorbance at 280 nm due to chromophore. Consequently, these two variables exhibited the most significant positive influence on the absorbance at 280 nm with respect to chromophore. On the contrary, an elevation in ultrasound exposure (A) and the chromatophore temperature (B) led to a reduction in absorbance at 280 nm, suggesting detrimental impacts on the isolation process.
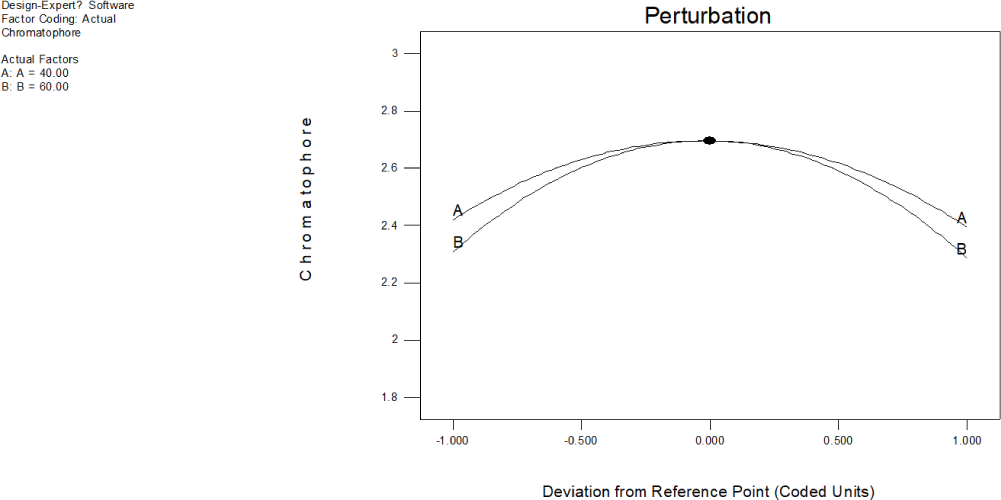
Linear effect of all factors at a point in a design space is compared using a perturbation plot of absorbance due to chromatophore at 280 nm for low (–1) and high (+1) levels of factor while keeping all other factors constant (Fig. 17). The perturbation plot of ommochrome illustrates how its response changes when deviating from reference point (coded 0) indicating that both exposure to ultrasound and the temperature, significantly influenced ommochrome extraction. Steep slope of the plot clearly indicates that change in absorbance due to chromatophore at 280 nm is very sensitive to the changes in one factor while keeping all other factors constant. Perturbation plot unmistakably illustrates that absorbance was very sensitive to change of temperature of the solvent (B:B = 60.00) than the changes in the exposure to ultrasound (A:A = 40.00).
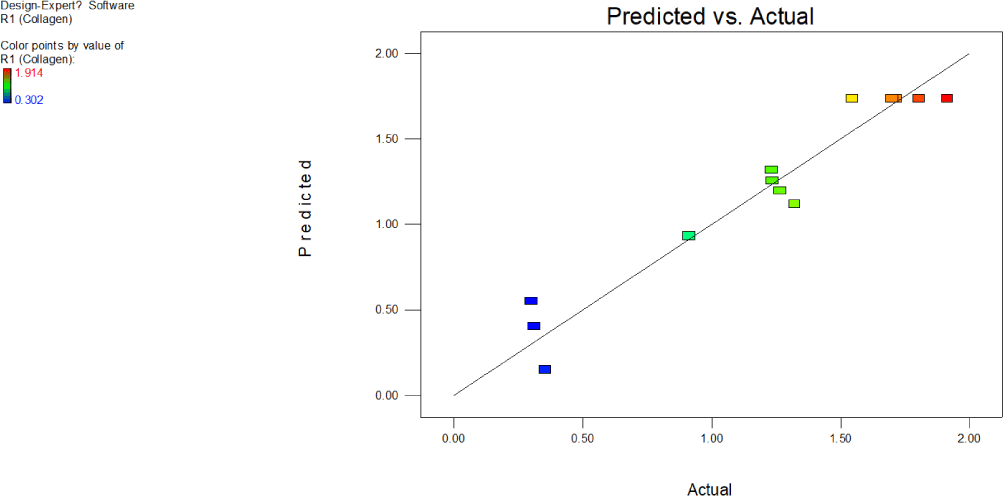
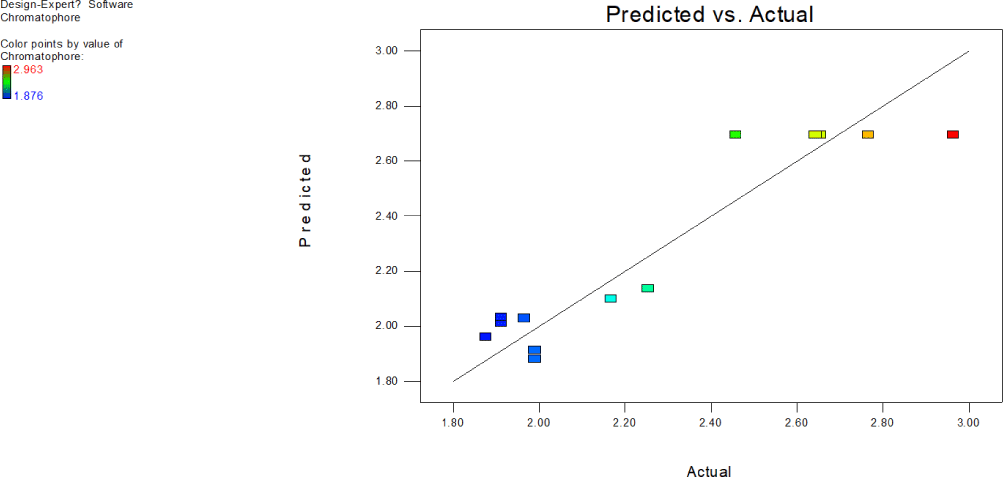
Predicted vale by the model for each data point is plotted against actual experimental value obtained to check how close closeness of the points (Fig. 18). The clearly shows that experimental values go positively with the predicted value. The increase in the absorbance values of ommochromes at 280 nm effected by exposure to ultrasound and temperature of the buffer are seen as not too far off from line scattered data points around the predicted line indicates model best fits. Based on the perturbation plot, conclusion can be drawn that both variables exposure to ultrasound and temperature of solvent significantly influenced chromatophore extraction.
RSM offers advantages by extracting experimental output from a limited number of actual experiments, analysing variable effects and their interactions, identifying factor levels that yield optimal responses, and determining optimal conditions for multiple responses. However, limitations of RSM are its limited ability to system with curvatures comprehensively. Increase in number of experiments with increase in number of variables, and reduction in prediction capability outside experimental domine, and limitations in explaining underlying mechanism even with its ability to describe outcomes under different conditions. Hence, ANN study is performed to make comparative capability of constructing experimental designs and predicting the efficiency of the response.
The Table 9 displays the R2 values for RSM, ANN, kNN and Random Forest models. This parameter quantifies how well RSM, ANN, kNN and Random Forest models fit the dataset. The R2 values explains how well RSM, ANN, kNN and Random Forest models can predict the response variable values. RSM, ANN, kNN and Random Forest models with attained R2 values of 0.93, 1.727, 1.734, and 1.642, respectively, fits well to the datasets. ANN model is more reliable and realistic since the predicted collagen value is closer to experimental results followed by kNN and Random Forest model.
Table 10 show optimized conditions through RSM with ML approaches for chromophore. Here, ANN showed highest R2 criteria score of 0.899, followed by Random Forest with 0.816, then RSM with 0.76 with and kNN with 0.371, all of which are above average. The R2 metric gauges the correlation between the predicted value and the target value (Kartal & Özveren, 2021), making it a frequently favoured parameter for assessing the model’s performance. It is important to highlight that the RSM, ANN, kNN and Random Forest models exhibit no increased difficulties in fitting. This is ascribed to the careful consideration given to the configuration selection process, wherein both the accuracy of training and testing, as well as the simplicity of the models, were considered.
This study successfully optimized the sequential ultrasound-assisted extraction of collagen and ommochrome from Indian squid (U. duvauceli) epidermal tissues. The squid, caught at 13.21505 latitude and 75.05525 longitude, measured 14–19 cm and weighed 86–98 g. From 1,000 g of whole squid, 103 ± 1.04 g of epidermal tissues were obtained, alongside mantle (452 ± 2.04 g), wings (21 ± 1.14 g), tentacle and other internal organs (424 ± 1.14 g). These epidermal tissues contained 22 g/100 g total protein and 3.95 g/100 g collagen.
For collagen extraction, optimal conditions, determined by CCD, were 30 minutes of ultrasound at 35°C, achieving a maximum absorbance of 1.74 ± 0.14 at 230 nm. This process yielded 2.76 ± 0.03 g of collagen per 100 g of epidermal tissues in the supernatant, with an additional 1.2 ± 0.02 g remaining in the pellets, resulting in an impressive total of 3.95 ± 0.04 g collagen per 100 g of initial epidermal tissue, indicating efficient recovery (Bavisetty et al., 2024; Sadowska & Sikorski, 1987).
Subsequently, the pigment sac-rich pellet fraction (containing 12.1 ± 0.15 g of pigment material per 100 g of initial epidermal tissue) was used for ommochrome extraction. CCD optimization revealed that 40 minutes of ultrasound at 60°C were ideal, yielding 1.24 ± 0.06 g of soluble ommochrome sample per 100 g of initial epidermal tissues in the supernatant, with a maximum absorbance of 2.69 ± 0.05 at 280 nm (Butenandt et al., 1957; Chan et al., 2019; Chen et al., 2023; Esparza-Espinoza et al., 2022).
Comparing predictive models, ANN outperformed OFAT, RSM, kNN, and Random Forest, demonstrating the highest coefficient of determination (R2) for accurately predicting extraction outcomes.
Conclusion
OFAT, RSM and ML (ANN, kNN and Random Forest models) approach were adopted to identify significant factors, perform CCD for sequential extraction of collagen and ommochrome from Indian squid skin. The selection of parameters and their corresponding levels during the initial OFAT experiments played a crucial role in the CCD of experiments for the sequential extraction of both collagen and ommochrome. RSM approach predicted optimum conditions for the maximum sequential extraction of collagen and ommochrome from Indian squid skin. CCD data were subjected to train the ML models for predicting reliability of experimental value of collagen and ommochrome from Indian squid skin. The optimal conditions for isolating collagen from chromatophore involve a 30.45 min exposure to ultrasound at 40 kHz, with a temperature of 38.8°C, and a buffer of pH of 2. Ommochrome extraction reached its peak efficiency when subjected to ultrasound at a frequency of 40 kHz for 39.77 min, utilizing HCl-MeOH with a pH of 2 at a temperature of 59.86°C. The current study demonstrates that ANN exhibits superior accuracy in predicting the sequential extraction of collagen and ommochrome from Indian squid skin compared to RSM, kNN and Random Forest models.